Summary
Part of the Genome in a Bottle Consortium hosted by NIST dedicated to comprehensive characterization of benchmark cancer genomes. Sign up for General GIAB and Analysis Team email lists.
Updates for May 2026:
- Our manuscript about the complete assembly of the HG008 pancreatic cancer genome, as well as somatic variant benchmarks from this genome is complete. The pre-print is available on bioRxiv ; "A complete human pancreatic cancer genome" https://doi.org/10.64898/2026.05.01.722316
- V0.5 HG008-T Draft Somatic Structural Variant and Copy Number Variant Benchmarks available on the GIAB FTP site. README describes the various benchmark files and how to use SV benchmarks with truvari for benchmarking. This is our first draft benchmark for subclonal SVs and we have refined instructions for benchmarking, so we welcome feedback about these: https://ftp-trace.ncbi.nlm.nih.gov/ReferenceSamples/giab/data_somatic/HG008/Liss_lab/analysis/NIST_HG008-T_somatic-stvar-CNV_DraftBenchmark_V0.5-20260318/
- V0.3 HG008-T Draft Clonal/Truncal Somatic Small Variant Benchmark. This is our first public somatic small variant benchmark, which includes many challenging variants that modify germline variants, particularly in homopolymers and tandem repeats. We welcome your feedback about best practices for benchmarking these: https://ftp-trace.ncbi.nlm.nih.gov/ReferenceSamples/giab/data_somatic/HG008/Liss_lab/analysis/NIST_HG008-T_somatic-smvar_DraftBenchmark_V0.3-20260425/
- New data for HG008 continues to arrive, including sequencing and karyotyping for several passages from 2 to 100 of the NIST HG008-T cell line: https://ftp-trace.ncbi.nlm.nih.gov/ReferenceSamples/giab/data_somatic/HG008/NIST/HG008-T_bulk/. Initial short read WGS of 8 HG008-T single cell clonal cell lines is available under https://ftp-trace.ncbi.nlm.nih.gov/ReferenceSamples/giab/data_somatic/HG008/NIST/HG008-T_clones/. We also have new data types in the data manifest from Dovetail Hi-C, 10x Genomics single cell ATAC+RNA-seq, MissionBio single cell targeted DNA, and PacBio Revio SPRQ and Vega WGS. Let us know if you are interested in helping analyze these data.
- We have initial sequencing of a second broadly-consented tumor cell line (HG009-T) and several clonal cell lines. This cell line is from a PDAC liver metastasis. We are currently attempting immortalization of a matched normal HG009 T cell line.
Interested in collaborating with us? Contact justin.zook [at] nist.gov (Justin Zook).
View the GIAB FAQ here.
Description

Goals:
This project is an extension of the Genome in a Bottle Consortium to develop the technical infrastructure (reference standards, reference methods, and reference data) to enable translation of cancer genome sequencing to clinical practice and innovations in technologies. The priority of GIAB is comprehensive characterization of human genomes for use in benchmarking, including analytical validation and technology development, optimization, and demonstration.
Reference Samples:
NIST has been collaborating with Andrew Liss at MGH to develop new tumor cell lines with paired normal samples that are explicitly consented for fully public dissemination of genomic data and cell lines. The first tumor cell line (HG008-T) is from a pancreatic ductal adenocarcinoma, for which we have paired normal pancreatic (HG008-N-P) and duodenal tissue (HG008-N-D) for sequencing, but no normal cell line. We currently are collecting extensive genomic data described below, and are working towards making these cell lines available in public repositories. We plan to have another pancreatic tumor cell line with a paired normal cell line in the near future, but these are still under development. We also welcome additional collaborations for tumor and normal cell line pairs that are explicitly consented for fully public dissemination of genomic data and cell lines.
Benchmark (or "High-confidence") Variant Calls and Regions:
We are working with the GIAB community to develop benchmark variants for the tumor and normal samples, using assembly-based and mapping-based approaches. We welcome collaborations in this new project.
Whole Genome Scale Data:
Starting in the Fall 2023, we began collecting a diverse set of whole genome scale measurements for the GIAB HG008 samples (Figure 1). We are making the data public, without embargo, as we collect them. The data being collected is described in Table 1. We welcome collaborations to analyze these data.
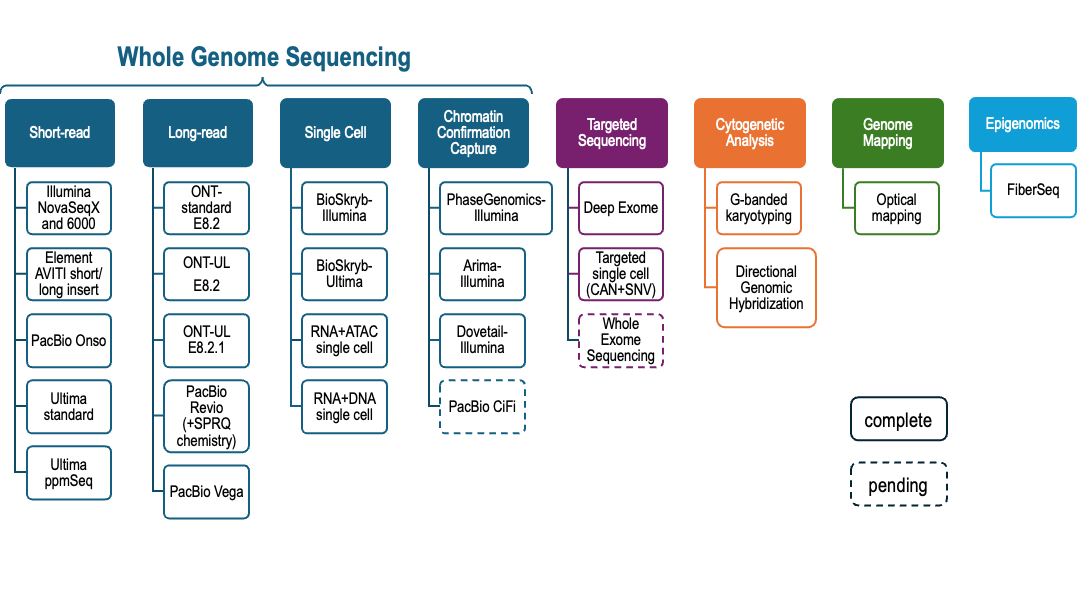
Figure 1 HG008 whole genome scale measurement technologies
Data Access:
These contributed data can be accessed through the public GIAB FTP as it becomes available. Data can also be browsed through 42basepairs , which allows for high level exploration and preview of the sequencing data.
For navigating the available data, we provide the Cancer GIAB Data Manifest, which allows for exploration of the tumor and normal data currently available on the FTP. If you are interested in exploring the manifest, you can create a filter view by 1) selecting the entire spreadsheet 2) Data → filter views → create new filter view. Please note tumor data collected from year 1 (2022) is from a prior passage of tumor cells and is emphasized in RED. Most of the tumor data being collected is from a single batch of tumor cells known as 0823p23. Please take these passages into consideration when choosing tumor datasets you are interested in exploring.
Table 1 Available datasets for GIAB HG008 tumor and normal samples
HG008 datasets that are available on the GIAB FTP have been QC'd and are noted by estimated coverage and read lengths. The Dataset ID corresponds to those in the Cancer GIAB Data Manifest. Coverage estimates reflect the expected coverage of diploid regions of the normal and tumor samples, assuming no whole genome duplication has occurred. Tumor data comes from either the bulk cell line or clones cultured from printed single cells from the bulk cell line. Note that a majority of tumor bulk cell line data comes from a large batch of cells (batch 0823p23), but some data are from other passages of the cell line at NIST and MGH as noted in the table. Clone passaging notes the passage of the bulk cell line the clone derives from plus the number of passages for the clone. This table is updated as data are received, last update 2026-05-04.
Cancer GIAB Publications:
- The manuscript describing extensive genomic data for the HG008 tumor/normal pair is now published in Scientific Data https://www.doi.org/10.1038/s41597-025-05438-2
- Our manuscript about the complete assembly of the HG008 pancreatic cancer genome, as well as somatic variant benchmarks from this genome is complete. The pre-print is available on bioRxiv ; "A complete human pancreatic cancer genome" https://doi.org/10.64898/2026.05.01.722316
Research Opportunities:
NIST-NRC Postdoctoral Fellowship: 2-year fellowship at NIST, U.S. citizens only, ~$82,000 salary plus benefits, application deadlines are Feb. 1 and Aug. 1, including up to 10 page research proposal. Contact Justin Zook if you are interested in writing a proposal on a genomics research project. We have opportunities posted for metrology in Cancer Genomics, Diploid Assembly, Epigenomics and Transcriptomics, Biological Data Science/Machine Learning, and Precision Medicine.

