Summary
The National Institute of Standards and Technology Genome Editing Consortium addresses the measurements and standards needed to increase confidence and lower the risk of utilizing genome editing technologies in research and commercial products. This consortium is currently active. Please see below for more detailed information on current activities.
MEMBER BENEFITS
- Access to a neutral forum for addressing pre-competitive needs
- Participation in the development of experimental benchmarks, guidelines, and terminology
- Access to tools developed by the consortium ahead of public release
- Institutional representation on consortium steering committee
BECOME A MEMBER
- Annual fee of $20,000 or in-kind support of equivalent value
- Participants will sign a Cooperative Research and Development Agreement
- Complete Letter of Interest Form
Notice of NIST’s Genome Editing Consortium Establishment
Notice of NIST’s Genome Editing Consortium Extension
PAST EVENTS
- 2025 Second Series of Workshops on Measurements and Standards for Advanced Therapy
- 2023 NIST-FDA Workshops on Measurements and Standards for Advanced Therapy - Workshop Report
- 2019 NIST Genome Editing Workshop
- 2018 NIST-FDA Genome Editing Workshop
- 2016 NIST Genome Editing Standards Workshop - Workshop Report
Description

Targeted genome editing, a method used to alter the DNA of living cells at desired locations, is poised to revolutionize science and medicine. To fight diseases, novel genome edited therapeutics, including those for use in regenerative medicine and infectious diseases, are being developed. Many commercial applications including agriculture and chemical production, are also leveraging this technology. Whether genome editing will be used in healthcare, agriculture, or basic research, robust quantitative measurements are needed to enable high confidence characterization of DNA alterations. NIST has brought together experts across the genome editing field including stakeholders in industry, academia, and government to assess their measurement needs. These discussions have identified common pre-competitive measurements and standards needed to establish greater confidence in the characterization of genome editing outputs. The NIST-led Genome Editing Consortium has been established to address these needs.
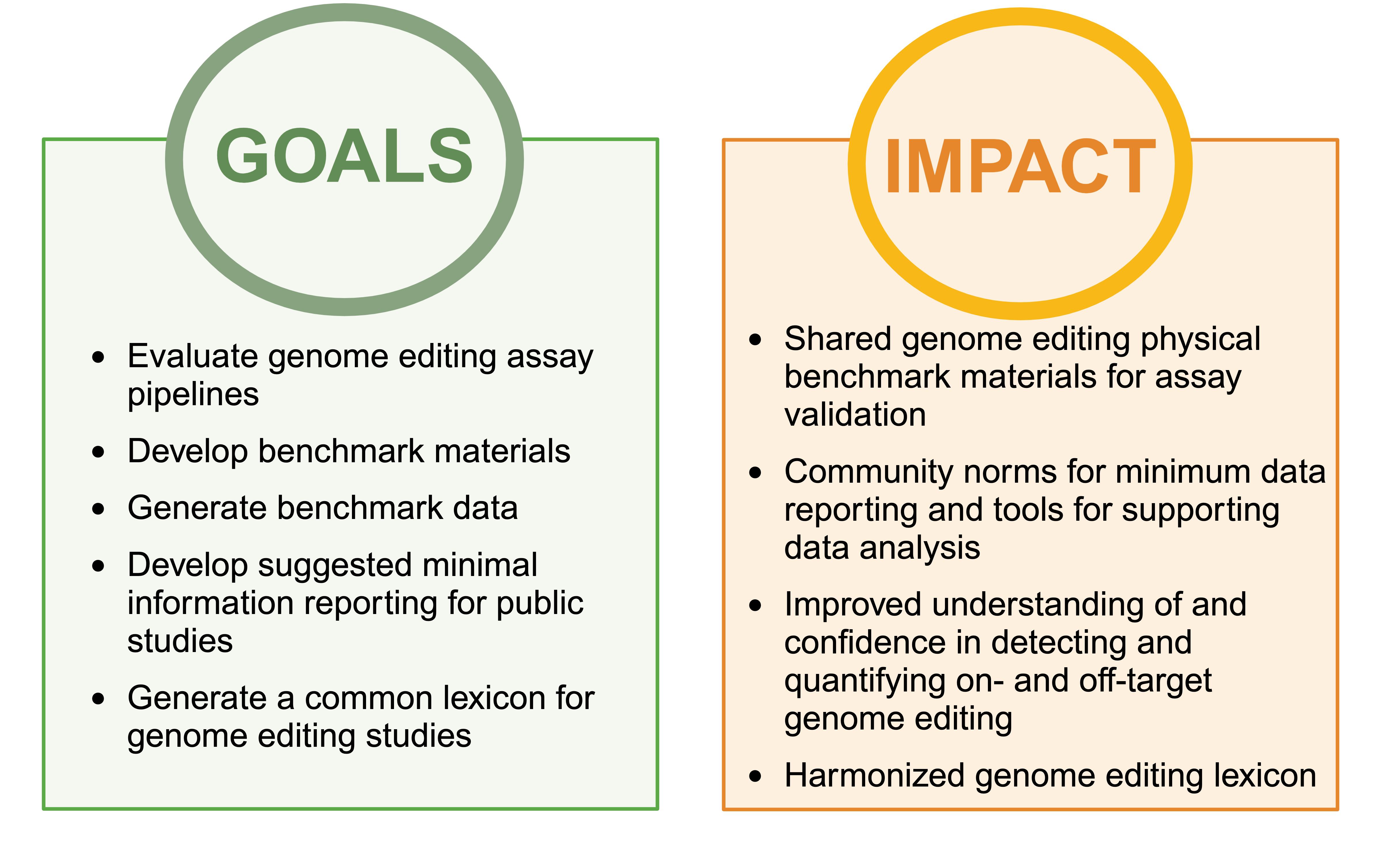

CONSORTIUM ORGANIZATION
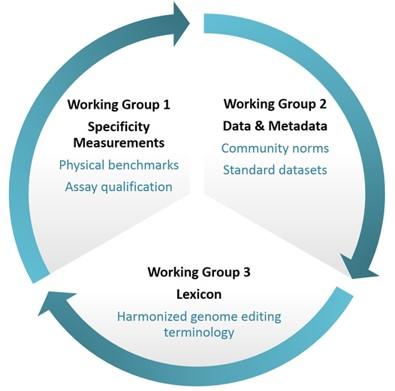
The consortium is composed of three primary working groups with the following responsibilities:
Working Group 1: Specificity Measurements
- Design, generate, and evaluate cell and DNA based control materials and test via interlab analysis. Control materials will be used as benchmarks for assessing next-generation sequencing (NGS) pipelines and other platforms intended to identify induced genome editing events.
- Design and conduct controlled evaluations of existing assays for quantifying on- and off-target genome editing, with a robust and optimal experimental design aimed at assessing the sources of variability, repeatability, and reproducibility within an assay.
Subgroups - NEW!
- Qualification of Off-Target Assays: Identify sources of variability and develop consensus approaches to qualifying off-target assays (including potential interlaboratory studies or control materials)
- Quality of Genome Editing Components: Identify concepts/information, approaches/assays, and potential controls for assessing quality of genome editing components
Working Group 2: Data and Metadata
- Identify community norms for data formats and tools for benchmarking data analysis including in silico and experimental data sets.
- Determine the type of metadata that would be needed to be shared, housed, and interrogated from genome editing experiments.
- Identify terms and related definitions to form a common genome editing community lexicon.
CONSORTIUM MODEL
- Convenes industry, academia, and government to identify and address measurement and standards needs across the genome editing field
- Enables members to work with NIST to develop measurement solutions and standards
- Leverages NIST expertise in measurement science, standards development, reference materials, technology development, and basic research
- Collaborates with related programs at other federal agencies
WHY NIST?
- Cross-disciplinary expertise in engineering, and the physical, information, chemical, and biological sciences
- As a non-regulatory agency of the U.S. Department of Commerce, NIST does not impose standards; standards are accepted by consensus
- Neutral convener for industry consortia, standards development organizations, federal labs, universities, public workshops, and interlaboratory comparability testing

