Summary
Flow cytometry and sorting bring new measurement capabilities to the field of microbiology, and microbial samples present unique challenges to traditional flow cytometry workflows. Flow cytometry uses scattered light and emitted fluorescence to quantify the properties of individual cells in a population. Many modern flow cytometers can detect microbe-sized particles (≈ 0.1 µm to 5 µm), and stakeholders are increasingly using flow cytometry to characterize microbial samples. Due to the method’s speed, multiplexing capability, and single cell resolution, there is great interest in using flow cytometry as a primary measurement or as validation for orthogonal characterization methods such as Coulter counting, colony forming unit assay, polymerase chain reaction, and sequencing. Excitingly, flow sorters enable physical separation of cell subpopulations for further analysis, for example isolating cells with distinct genome copy numbers.
Standardization of microbial flow cytometry methods is difficult due to aspects of microbial samples, including small cell sizes, irregular cell shapes, acellular environmental matrix, inconsistent response to fluorescent probes across cell types, and lack of microbe-specific fluorescent reagents. Our goal is to develop experimental and data analysis methods to advance stakeholders’ measurement assurance strategies. Application areas span microbial cell reference materials, biomanufacturing and biotechnology, biotherapeutics (“bugs as drugs”), pathogen detection, sterility testing, biosurveillance, and microbiome studies.
Description
Microbial flow cytometry capabilities support numerous ongoing projects, including:
- Microbial Cell Enumeration and Characterization: Establishing standards, measurement methods, and Reference Materials to enable stakeholders to assess accuracy, bias, and comparability across microbial enumeration measurements and microbial cell products.
- Human Gut Microbiome Reference Material (RM 8048): Whole stool reference material with measured homogeneity and stability and characterized for several measurands, including cell content via flow cytometry.
- Rapid Microbial Testing Methods Consortium: Characterization of reference and test materials to support Consortium activities such as microbial cell reference material quantification, method development, and interlaboratory studies.
- Highly characterized microbial materials used as measurement method inputs or as inter-sample controls (“spike-ins”).
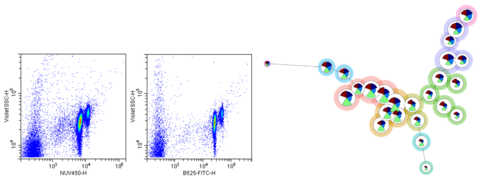
Figure 1: Left – Scatterplots showing the distribution of DNA intensity (x-axis) and violet side scatter (y-axis) as measured by two different fluorescent probes. Right – FlowSOM tree diagram representing potential particle subsets in a complex microbial community (background circle color) based on fluorescence measurements (intra-circle wedge color and magnitude).

