Summary
Currently, forensic DNA profiles consist of size measurements which are interpreted as the number of repeat units at STR (short tandem repeat) markers. Due to the decrease in cost of modern sequencing technology (known as next generation sequencing, or NGS), manufacturers are developing assays designed for forensic DNA applications. These new tests will allow forensic scientists to sequence STR markers, potentially resulting in increased ability to differentiate individuals in complex mixtures. Additionally, alternate marker types such as SNPs (single nucleotide polymorphisms) can be more easily integrated into casework laboratories, resulting in new capabilities such as ancestry or phenotype prediction in unsolved cases. The NIST Applied Genetics group supports this transition by beta testing assays, contributing to nomenclature and standards development discussions, and providing sequenced reference materials.
Description
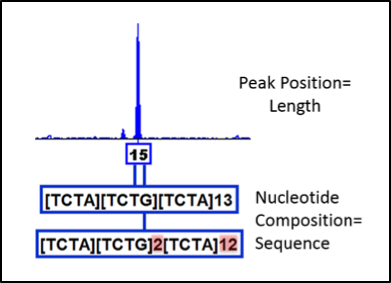
Forensic DNA profiles have historically been generated by Capillary Electrophoresis (CE), which results in length-based designations reported as the number of repeats at STR (short tandem repeat) markers. For example, the marker D3S1358 contains repeats of the four nucleotides “TCTA” or “TCTG”. When there are 15 total repeats, the designation is a 15 allele as shown in the example output. Differences in these length-based measurements between individuals are used to exclude individuals as potential donors of genetic material at crime scenes. However, there may be additional variation that is not captured with this method. By analyzing the DNA sequence comprising the peak, multiple versions of a 15 allele can be detected, as depicted below the peak. This example is from a single individual, but the potential for using sequence data to distinguish overlapping alleles in mixtures of multiple individuals can be readily realized. However, it is important to fundamentally understand this technology prior to forensic casework applications. Considerations such as limit of detection, artifact characterization, sequence-based allele frequencies, bioinformatics, and nomenclature, are being investigated at NIST.
NGS may also encourage adoption of alternate marker types by the forensic DNA community. Single nucleotide polymorphisms (SNPs) are the most common type of variation in the genome. As the name implies, they consist of a single base change—where one individual may have an “A” (abbreviation for the nucleotide Adenine), another individual has a “G” for Guanine. Like STRs, SNPs may also be used to identify individuals. In particular, multiple SNPs in a single DNA sequence (dubbed microhaplotype) can rival an STR locus in discrimination capacity and may be less challenging to sequence and analyze. Some SNPs are useful for bio-geographical ancestry (BGA) prediction of an unknown crime scene sample, and other SNPs may be used to predict externally visible characteristics (EVC) such as eye color. The ability to predict BGA or EVCs present new challenges to the forensic laboratory, and NIST scientists are evaluating prediction capabilities across assays, models, and populations.

