Integrated structural biology for predicting and quantifying nucleic acid structure and dynamics
Summary
Ribonucleic acid (RNA) dynamics include the time-dependent structural changes that take place during the life cycle of RNA molecules. Deciphering the dynamics of RNA is crucial to develop nucleic acid (NA) biotherapeutics, create synthetic biological circuits, and to understand gene regulation. This project synergistically combines multiple biophysical techniques for characterizing RNA structure and dynamics, including NMR and SAXS, with molecular dynamics simulations as the linchpin. The project progresses along three major directions:
- Comparative assessment and improvement of atomic force field potentials in refining RNA structures with solution state data
- Structure/ensemble prediction for highly dynamic motifs/hairpin loops (e.g. COVID SL2&3, 2GBH loop)
- Assessment and improvement of AI-based RNA structure prediction with experimental data.
Description
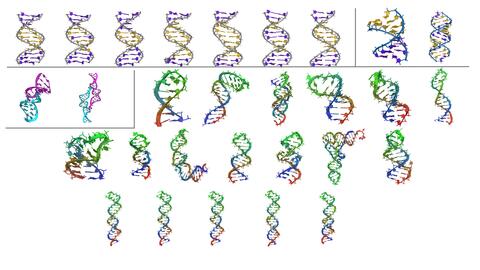
- Comparative assessment and improvement of atomic potentials. NMR is a primary structure determination method for short RNAs (100 nucleotides or less). Many types of NMR experiments can be performed, yielding data with varying levels of precision in defining structure: nuclear Overhauser effect distances (NOEs), chemical shifts, J-coupling constants (Jcc’s), and residual dipolar couplings (RDCs). A force field must be used to interpret these signals and convert them into a structure. Our prior work evaluated the ability of molecular dynamics force fields to reproduce experimental data for an A-form duplex (2GBH), showing that unrestrained molecular dynamics simulations reproduces experimental solution state RDC data. Additionally, the effect of water models on the ensemble fitting of RDC data was examined. Loop structure and dynamics for SARS-CoV-2 5’UTR viral RNA stem loops were evaluated, and model helices were predicted. We will build on these studies by investigating the impact of force field modifications on the fidelity of simulated structural ensembles with respect to the accessible experimental data.
Structure/ensemble prediction. A challenge in defining RNA dynamics includes a shift in defining structure from a single entity to an ensemble of structures with respective weights, that all contribute to a measured signal. This challenge is much more similar to intrinsically disordered peptides, where the single stranded nature and formal charge, as well as high number of rotatable bonds in comparison to proteins, increases the potential conformational space accessible to a nucleic acid. Additionally, ensemble sampling methods must be developed and used to rigorously search this space. We used an RNA hexamer to understand the interplay between different types of solution state data, SAXS and NMR, and prediction methods, all atom molecular dynamics and physics-based/template structure predictors. Our work has helped integration of multiple biophysical methods. These studies will be carried over to a select set of flexible RNA constructs that have so far resisted structural interpretation despite the abundance of available experimental data.
Machine learning for RNA structure prediction. A limiting problem in the application of current structure prediction methods to RNA, in particular, is the lack of training data available. While proteins have orders of magnitude more structures available on which to train ML predictors, RNA is primarily limited to large ribosomal subunit structures or double stranded structures. This, in turn, limits the predictions to base pared vs. unpaired levels of information. We maintain a database of highly curated structures with the accompanying high-precision NMR RDC data, which can be used to benchmark structure prediction algorithms. These structures span multiple important conformational states and include duplex and triplex helices, tetraloops and longer loops, alternative base pairings, bulges, and motif structures like T-RNA and the UUCG tetraloop.
Data: https://github.com/usnistgov/AI-Structure-Prediction
Publication: https://pubs.acs.org/doi/full/10.1021/acs.jcim.5c00245.
Publications/Presentations
Bergonzo, C. and Grishaev, A. Accuracy of MD solvent models in RNA structure refinement assessed via liquid-crystal NMR and spin relaxation data. J. Struct. Biol. 2019, 10.1016/j.jsb.2019.07.001.
Bergonzo, C. and Grishaev, A. Maximizing Accuracy of RNA structure in refinement against residual dipolar couplings. J. Biol. NMR. 2019, 73, 117-139. 10.1007/s10858-019-00236-6
Olenginski, L., Kasprzak, W.K., Bergonzo, C., Shapiro, B.A., Dayie, T.K. Conformational dynamics of the hepatitis B virus pre-genomic RNA on multiple time scales: implications for viral replication. 2022, J. Mol. Bio., 167633. 10.1016/j.jmb.2022.167633
Bergonzo, C.*, Grishaev, A., Bottaro, S. Conformational heterogeneity of UCAAUC RNA oligonucleotide from molecular dynamics simulations, SAXS, and NMR experiments. RNA. 28, 7, 937-946. 10.1261/rna.078888.121
Bergonzo, C.* and Szakal, A.L. Using all-atom potentials to refine RNA structure predictions of SARS-CoV-2 stem loops. Int. J. Mol. Sci. 2020, 21, 17, 6188. 10.3390/ijms21176188
Bergonzo, C.* and Grishaev, A. 2024. Critical assessment of RNA and DNA structure predictions via Artificial Intelligence: the imitation game. J. Chem. Inf. Model. 65, 7, 3544-3554. **Awarded “Editor’s Choice”** 10.1021/acs.jcim.5c00245
“RNA dynamics” RNA Dynamics Workshop, Telluride CO July 2025
“Critical assessment of RNA and DNA structure sing MD, ML, and experiments: the rise of the machines” IBBR Structural Biology Day Invited Talk, May 2025
“Critical assessment of RNA and DNA structure predictions from AI: The imitation game” Co-presented with Alex Grishaev, NIST Group Meeting October 2024
“Computational Prediction of RNA HOS” OPTCongress March 2022
“Refining Predictive Models of RNA Therapeutics” RNA Dynamics, Telluride CO July 2021

