The workshop will bring together multiple research and development (R&D) communities focusing on big image analyses in computer cloud environments. Such analyses are frequently supported by implementing web client-server systems executing a wide spectrum of algorithms designed to extract image-based measurements, and perform image classification, object detection, object registration, object tracking, and object recognition. The purpose of this workshop is to discuss the bio-medical and bio-materials science application needs for big image analysis solutions, current open-source technical solutions, and community-wide R&D interests in defining inter-operable algorithmic plugins for web client-server systems designed for big image analyses.
Background:
There is an increasing interest in enabling discoveries from high-throughput and high content microscopy imaging of biological specimens and bio-material structures under a variety of conditions. As automated imaging across multiple dimensions increases its throughput to thousands of images per hour, the computational infrastructure for handling the images has become a major bottleneck. The bottleneck presents challenges that range from transferring data, storing and archiving, annotating, quantifying, and visualizing, to the mechanisms for applying the latest machine learning and artificial intelligence models by non-computational experts from a variety of application domains. These challenges arise due to big image data, complex phenomena to model, and non-trivial computational scalability that accommodates advanced hardware and cutting-edge algorithms. Furthermore, the challenges are amplified by the need to engage a broad community of experts in analyzing complex image content and the need to reproduce discoveries based on image measurements and any decisions derived from these measurements. Such measurements, discoveries and decisions are critical for biological and bio-materials science applications, for instance, quality assurance of stem cell therapies, design of cancer treatments, high throughput screening in drug discovery, and vaccine discoveries from atomic resolution structures of viruses and protein complexes.
Existing solutions: To overcome the aforementioned challenges, several research institutions have prototyped web-based systems in order to facilitate access to large image databases and to high performance computing (HPC) and cloud hardware resources. The existing web-based prototype solutions leverage a variety of web technologies on the client side and a spectrum of databases, scientific computational workflow engines, and communication protocols on the server side in order to hide the infrastructure complexity from the domain application experts and make them more productive in conducting research. While the web-based solutions deliver infrastructure capabilities, their capabilities for processing large images remain limited to the computational tools provided by each development team because the development of new tools is web solution specific and a definition of an inter-operable web computational plugin does not exist.
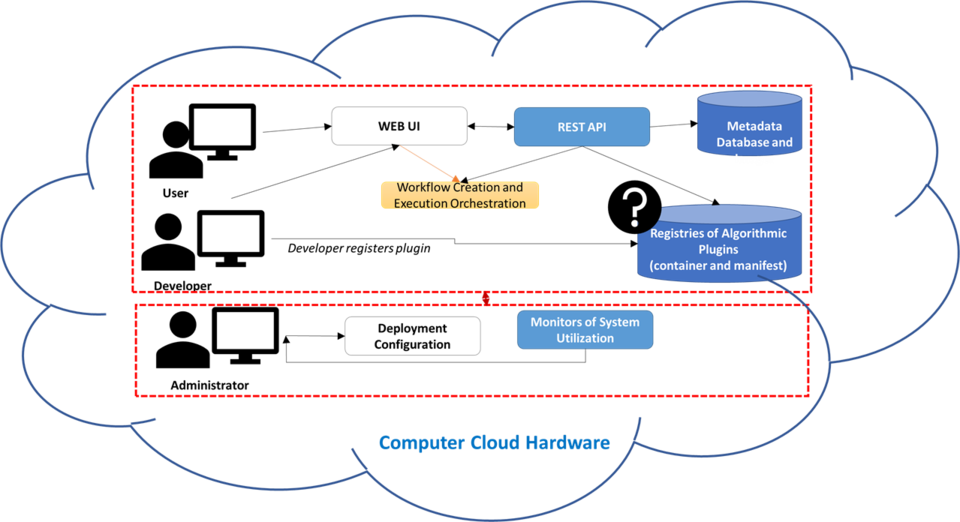
With the increasing popularity of software containers as standardized units for deployment, there is an opportunity for the communities working with large microscopy images to discuss creating inter-operable web computational plugins. These web computational plugins consist of software containers and web user interface (UI) description files to enter parameters needed for the software execution. Each container packages code with all its dependencies and has an entry point for running the computation in any computing environment. Each UI description file contains metadata about the plugin container and the computation parameters. This description file is intended for generating web UI for entering parameters dynamically.
Scope, Goals, and Topics:
The workshop scope will touch on the technical topics of big image data management, software containerization, software execution on advanced hardware architectures, container-based workflow management, dashboards for monitoring computations, web technologies for dynamic content creation, web-based visualization of large images, delivery of provenance information, and web plugins for image annotation creation. The main goal for the workshop is to establish a community consensus on creating inter-operable web computational plugins that can be chained into scientific workflows/pipelines and executed over large image collections regardless of the cloud infrastructure components.
Vested interests in the workshop topics: Multiple government agencies, non-profit research institutions and commercial entities have invested large amounts of resources into building software for big data analyses and setting up hardware infrastructure for cloud computing and high-performance computing. To improve the utilization of all investments, private and public sectors might benefit from inter-operability of cloud deployable algorithmic plugins. Specifically, improving the reuse and execution of open-source and close-source algorithms applied to big image analyses is of interest to bio-medical and bio-materials science stakeholders as the domain problems are complex and need very large number of image observations. We plan to facilitate discussions by forming working groups focusing on:
- Containerization of execution code
- Data storage and access interfaces for object-, block- and file-level storage
- Inter-operability requirements of workflow engines for running containerized plugins
- Standard packaging of web UI modules
- Security of container-based distribution
Agenda
Overview
The two-day workshop is divided into information exchanges (day 1) and technical discussions (day 2). During the information exchange day, speakers will deliver
- Category 1 Presentations
- Introduce bio-medical application needs for big Image analysis
- Introduce bio-materials science application needs for big image analysis
- Category 2 Presentations
- Describe the current open-source technical solutions
- Describe the current commercial technical solutions
- Category 3 Presentations
- Outline the funding mechanisms behind the R&D efforts
During the technical discussions, attendees will identify common interfaces for web deployable algorithmic plugins across existing open-source web client-server systems in terms of the following:
- Category 1 Coding Working Group
- Describe expected user inputs and algorithmic outputs for creating dynamic web applications
- Category 2 Packaging Working Group
- Discuss universal packaging of image analysis algorithms for cloud deployment
- Category 3 Workflow Working Group
- Presenting a workflow of chained plugins for executing the plugin computations
Schedule Day 1 - (All session during Day 1 will be in Lecture Room D)
| Time | Event | ||
| 8:30am | Registration and continental Breakast | ||
| 09:15 | - | 0930 | Opening remarks – Goals for day and meeting |
| 09:30 | - | 11:30 | Presentations focused on the-state-of-the-art and future needs in large size microscopy image analyses
|
| 11:30 | - | 12:30 | Lunch (on your own) |
| 12:30 | - | 02:30 | Presentations focused on funded activities and possible funding opportunities
|
| 02:30 | - | 02:45 | Coffee break |
| 02:45 | - | 04:45 | Presentations focused on existing client-server solutions
|
| 04:45 | - | 05:30 | Closing Remarks – Goals for Next Day |
Day 2
| Time | Event | ||
| 8:30 | Registration and Continental Breakfast | ||
| 09:15 | - | 0930 | Summary of Day 1 - (Lecture Room D) |
| 09:30 | - | 10:30 | Working groups: focus on code review of plugins in existing web client-server systems (*5 rooms as specified below) |
| 10:30 | - | 11:30 | Poster session – architectures and demonstrations of plugins in existing web client-server systems (Lecture Room D) |
| 11:30 | - | 12:30 | Lunch (on your own) |
| 12:30 | - | 02:30 | Working groups: focus on summarizing common characteristics of plugins in existing web client-server systems and on desirable characteristics of such plugins (*5 rooms as specified below) |
| 02:30 | - | 02:45 | Coffee break |
| 02:45 | - | 04:45 | Summaries from working groups and open discussions (Lecture Room D) |
| 04:45 | - | 05:30 | Closing Remarks and Next Steps (Lecture Room D) |
*Working Group Locations:
- Containerization of execution code (Lecture Room A)
- Data storage and access interfaces for object-, block- and file-level storage (Lecture Room B)
- Interoperability requirements of workflow engines for running containerized plugins (Lecture Room D)
- Standard packaging of web UI modules (Portrait Room)
- Security of container-based distribution (Heritage Room)
Lodging Information
Double Tree by Hilton
620 Perry Pkwy, Gaithersburg, MD 20877
The hotel provides a complimentary shuttle to and from NIST for those who booked under the meeting room block. If you book outside the meeting room block you will need to provide your own transportation.
Security Instructions
*Visitor Access Requirement:
For Non-US Citizens: Please have your valid passport for photo identification.
For US Permanent Residents: Please have your green card for photo identification.
For US Citizens: Please have your state-issued driver's license. Regarding Real-ID requirements, all states are in compliance or have an extension through October 2020.
NIST also accepts other forms of federally issued identification in lieu of a state-issued driver's license, such as a valid passport, passport card, DOD's Common Access Card (CAC), Veterans ID, Federal Agency HSPD-12 IDs, Military Dependents ID, Transportation Workers Identification Credential (TWIC), and TSA Trusted Traveler ID.

