Summary
Technologies to manipulate biological systems have far outpaced our ability to design the function of those systems in a predictable way. This is, in part, a consequence of the well-known complexity of life. Simplified biological systems offer increasingly accessible solutions that are reduced to their essential features, while remaining complex enough to be alive. This approach of simplifying a biological system for study can help identify the mechanisms and emergent properties underlying that system's full complexity and function. The Cellular Engineering Group at NIST has partnered with collaborators to introduce and characterize minimal cells as a potential enabling platform for fundamental research and industrial applications.
Description
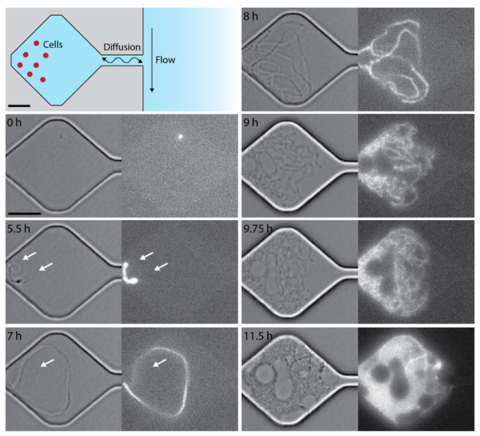
Dynamic cell division and growth in microfluidic chambers show diverse morphotypes emerging from a single, genomically minimized cell. Cells diffuse freely in a microfluidic chamber, which exchange nutrients and waste diffusively with a larger channel (scale bars: 5 µm).
Minimal cells are cells whose genome has been stripped of all but those genes essential for replication, in a process called "genome minimization." NIST and collaborators demonstrated genome minimization coupled to single cell imaging as a powerful approach to investigate the minimal genetic requirements for cell division and control of cell size and shape. Size and shape are fundamental aspects of cell physiology that consequently serve as an ideal initial test case for using genomically minimized cells to identify predictive relationships between genotype and phenotype. Designing whole genomes to achieve desired phenotypes across multiple scales, from single cells to populations, during whole genome engineering stands as a grand challenge in synthetic biology.
This collaboration links scientists at NIST to the Synthetic Biology Group at JCVI, headed by John Glass, James Pelletier formerly at the Center for Bits and Atoms at MIT working under Neil Gershenfeld, and imaging facilities at UCSD.

