CHRNS: Event mode software
Overview
With the improvements in hardware, sample environment, and new data packets/storage of events we require new software for the production, distribution, caching, saving, histograming, reducing, fitting, monitoring, and feedback to users/NICE.
- CHRNS Computing resources are available via ORCiD login within NIST
- DAVE and Mslice for MACS
- SASView for VSANS
- Refl1D for CANDOR
- Developing necessary web-based tools to perform all other tasks
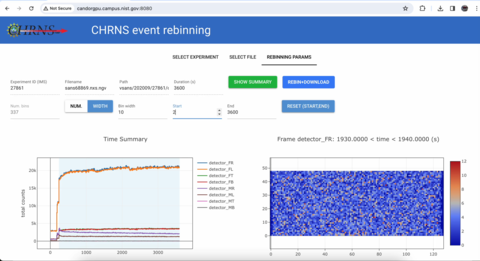
New rebinning API and web interface for event mode data
Built around the newest data management tools from the CHRNS-FAIR initiative, we have designed a web interface that allows inspection and rebinning (<1 second!) of time-stammped neutron data. The API will allow for automated data binning. The code is located here.
Refl1D/CANDOR data fitting
- Enabled remote (web-based) access to a 'live' fitting server for interactive fitting sessions, or static fit results for batch fits for the Refl1D analysis software, which is used for analysis of data from the CHRNS CANDOR instrument.
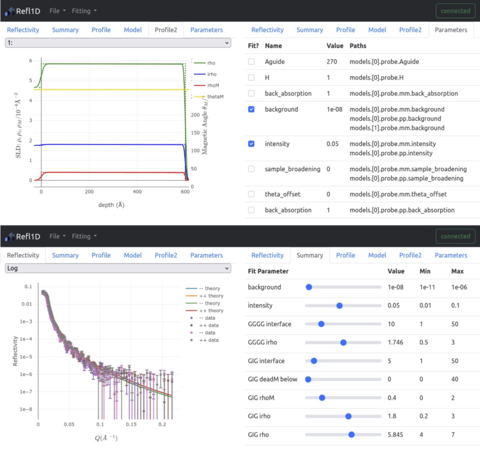
- The interface is detachable and re-attachable to the fitting session at any time
- Parameter error bars are determined with MCMC, correlations, fit history etc. visually represented
- Necessary for Autonomous Experiments
- Low-code fitting development:
- Simple web app lets users build physical models
- Compare model to data directly in-app
- Code is generated from the visual model representation and piped to Refl1D
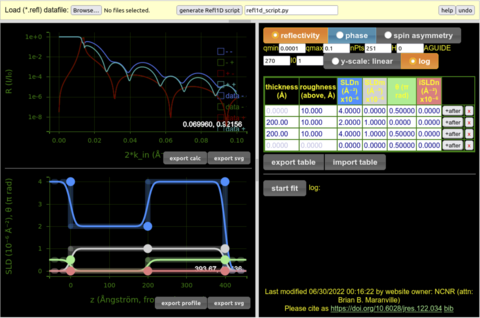
Experimental optimization of the neutron instrument, measurement, and sample to greatly reduce measurement times and increase data quality.
- Remote Execution of CANDOR experimental optimizations
- Jupyter Notebook data fitting using Refl1D
- Feature-complete experimental optimization for bio-materials
- C++ classes wrapped in Python for speed and compatibility
- Declarative model langue for use in Refl1D
- See section on CANDOR Fluids handling robot for 1) ‘immediate’ feedback to the user on the information content in a measurement 2) a mechanism for feeding results from these routines back into the NICE control software to be used in intelligent, automated determinations of how to efficiently complete the measurements.
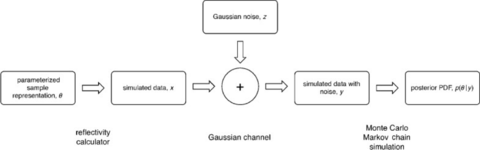
using Bayesian statistics and information theory
SASView/VSANS data fitting and modeling
- decoupled GUI from backend computations
- GPU/CPU multiprocessor support for fitting
- SasView: https://github.com/SasView/sasview
- SasModels: https://github.com/SasView/sasmodels
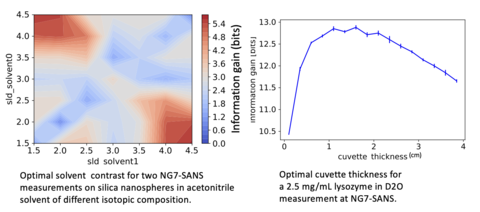
Intelligent experiment-design and experiment-planning tools using predictions of information content based on user-supplied models (bio/soft materials initially)
- Web-based Application : (login using ORCID inside NIST: https://charlotte.ncnr.nist.gov/jupyter)
- Remote Execution of SANS experimental optimizations
- Feature-complete experimental optimization
- lipid vesicle molecular model
- Any SASView model
- Monte Carlo Markov Chain fitting of SANS data
- Instrument resolution
- this is also needed to classifying SANS patterns with machine learning (in collaboration with nSOFT)
- We have implemented Autonomous Reflectivity measurments on CANDOR and once we gain experienence and user feedback, we will use that understanding to provid intelligent, automated determinations of how to efficiently complete VSANS measurements.The ‘immediate’ feedback to the user on the information content in a measurement and statistical error analysis for a given model(s) can be used to reduce the amount of overcounting on SANS. This does not require a time-resolved experiment to benefit from the capability.
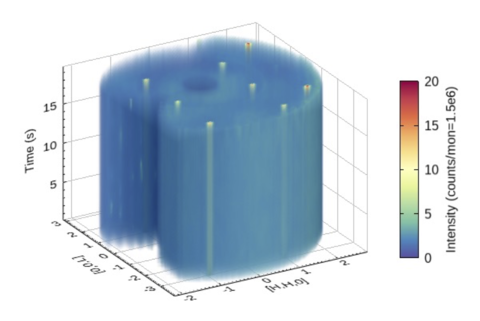
DAVE and Mslice
Used for reducing MACS data and has been adapted to visualize time-slice data, taking special care of the error bar due to low count rate of the event mode. (DAVE)
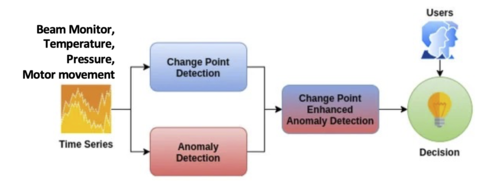
AI error monitoring
While NICE still has monitoring capabilities, for the instruments, we are looking how to implement AI/ML routines providing real-time reporting of unexpected experimental conditions, such as significant statistical data fluctuations or sample environment deviations. User alerts and automated instrument responses can be added. We have 2 possible unsupervised methodologies (that work in different scenarios - but not suitable for every experiment), and are working on generating labeled data (from past sample environment records) to work with Microsoft and implement a general monitoring.
Contacts
-
(301) 975-6034

