Taking Measure
Just a Standard Blog

We are all well aware of COVID-19, and by now most people have seen pictures of the spike protein that forms the “handshake” interaction between virus and host cells and is the basis of two new vaccines. The COVID-19 virus is made of RNA, which manufactures the spike protein and all the other proteins that allow it to survive. What if scientists could target the RNA in the virus before that manufacturing process even begins? That’s where my work centers, around COVID-19’s viral RNA.
Before the proteins that infect your cells can be built, the viral RNA, which contains the blueprints to produce proteins essential for viral replication, must be read by the ribosome, the place where proteins are put together within a cell. Parts of the viral RNA form flexible structures that regulate the ability to read it and create proteins from it. If we can develop drugs that interfere with these RNA structures forming, the virus can’t function. In my work at the National Institute of Standards and Technology (NIST), I use computer simulations to predict what these RNA structures look like and how they move to gain a better understanding of how they can be targeted by drugs.
RNA structures are tricky to predict compared with protein structures for a few reasons. We have 3D maps of many more proteins (about 40 times more!) from all types of organisms, so algorithms to predict RNA structures start out with much less information about what they might look like. This is because RNA can be difficult to work with experimentally — so there are fewer tools available, at a higher cost, and generating data from experiments takes a long time. Also, RNA can be very flexible (aka “dynamic”), adding complexity to the prediction. Frequently, the same piece of RNA can form multiple structures, and we take them all into account to create an ensemble, a group viewed as a whole rather than as single, individual parts.
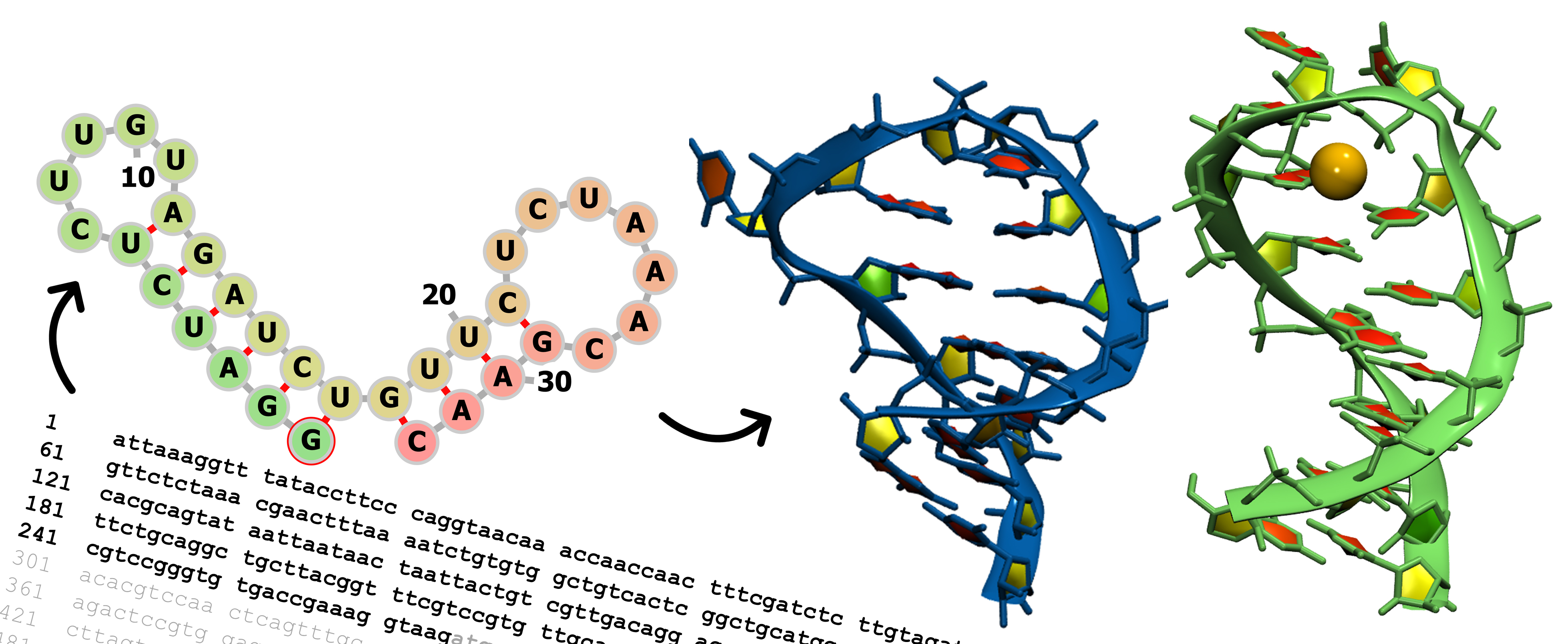
We are looking at RNA in a section of the COVID-19 viral genome that sets up translation, which is the process of reading the RNA to create the proteins the virus needs to survive. Specifically, two short regions called Stem Loop 2 and Stem Loop 3 (SL2 and SL3) contain important parts of the RNA that interact with other parts of the RNA to control the manufacture (expression) of proteins. They are called stem loops because the genetic sequence — repeats of the “letters” C, U, A, G — pair up C-G and A-U at the beginning and end of the sequence into a helix to form a ladder-like stem, while the middle part of the remaining sequence is unpaired in a loop. The RNA in SL2 has the same genetic sequence as the SL2 in the coronavirus SARS-CoV-1, the virus that caused severe acute respiratory syndrome (SARS) in the early 2000s.
So we hypothesized that the SL2 in the COVID-19 virus must adopt the same 3D structure found in the earlier SARS virus. However, computational predictions, which try to match sequences to parts of already determined RNA structures, generated 3D structures very different from the reported SARS-CoV-1 SL2 RNA. We wanted to find out if that would change using more advanced computational prediction.
Using all-atom molecular dynamics simulations, where we explicitly model the RNA, water and ions that would be present in the cellular environment, we find that the RNA rearranges quickly to match the RNA loop structure from SARS-CoV-1 with the same sequence. This shows that the all-atom molecular dynamics can adjust previous rough predictions that might not show the fine details of structure and resolve the dynamics of the RNA. And this means we can use it to predict details for something that we don’t have any other information on — a blind prediction.
For instance, SL3 is another short piece of the viral RNA that we think forms a loop. In many coronavirus genomes, there is something called a transcription regulatory leader sequence here. This transcription regulatory leader sequence helps to control protein expression. Some viruses have this piece of RNA unstructured, or flexible and able to adopt many different shapes, while other viruses, such as the one that causes COVID-19, are predicted to have this part of the RNA structured, or rigid and resistant to taking on different shapes. This RNA structure would also need to be easily disrupted for it to do its job and interact with other parts of RNA — making predicting its dynamics important if we are going to try and change them!
Simulations of SL3 show us that it is very flexible and adopts many different structures. Every so often, a potassium ion binds to SL3 and stabilizes a particular structure, creating a scaffold so the region we’re looking at can be recognized by RNA from further away. This allows for the RNA in this region to have enough structure to trigger reading, while making it easy enough to eliminate structure so the reading can smoothly progress. You can imagine it as a draw bridge, which needs to be down for cars to pass over it and up for boats to pass under it — like the SL3 RNA, both orientations are essential to its job.
We know that the computer simulations that we use result in models that are accurate because they are close to structures that we have actually seen in lab experiments. By having confidence in the computer simulation methods, we can extend them to other parts of the RNA. The SL3 computational prediction links the RNA structure to known function for how transcription is controlled. Using molecular dynamics to link and predict structure and function is the goal of these computational methods.
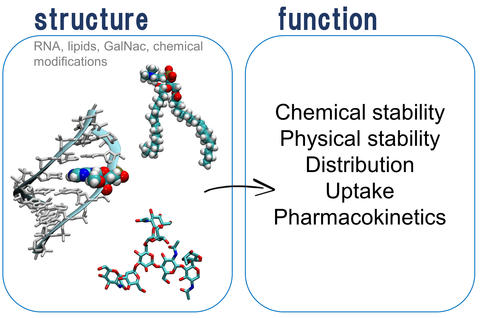
Predicting RNA structure is also important for developing drugs and vaccines where the RNA is itself the “active ingredient,” as in the Pfizer and Moderna COVID-19 vaccines. In these vaccines, the RNA needs to interact with other “ingredients” to come together in a formulation that can get the RNA into cells in the right amount of time, allowing its code to be read by cellular machinery, while remaining stable in vials in the clinic at reasonable temperatures. By understanding structure and function, we can engineer stability into drug products, optimizing for downstream manufacturing concerns such as avoiding extremely cold storage temperatures, for example.
We are using computer simulations and all-atom molecular dynamics to predict how these pieces interact and how we can change the ingredients to help make stable vaccines. This expands work done under the NIST Biomanufacturing Initiative, a program that to date has largely focused on measurements and standards to support development of protein-based drugs, to RNA-based drug platforms. Given the technical challenges, cost and time required for exploratory experimental work in the development of RNA-based drug platforms, application of fast computational algorithms to perform the biophysical characterization that is central to our work at NIST can be used to save stakeholders time and money, and to help expedite bringing these life-saving drugs to the public.
About the author
Related Posts
Comments
CONGRATULATIONS¡
Hi Dr. Bergonzo,
Your article is thought-provoking. I believe your line of inquiry would be very helpful if successful eventually.
I am also interested in Biomolecular Structure and Function, but am a novice in this field. I recently retired as Professor of Aerospace Engineering Sciences at the University of Colorado, Boulder, and hence have some time for other topics of curiosity. However my background is fluid flows, rockets and aircraft engines, and nothing in RNA structure etc. Can you please suggest a book or journal article(s) (yours, hopefully) to get started? Also, is the RNA sequence of COVID19 virus available in digital format to play with? I am fundamentally interested in the helical and loop structures of RNA. I would like to better understand how a helix or a loop is formed or why one is preferred over the other. Thanks in advance. My e-mail is [email protected].
Thank you for your compliment and comment!
You can find the complete primary sequence of the virus deposited in the NCBI database, and is linked here: https://www.ncbi.nlm.nih.gov/nuccore/NC_045512
You can find the proposed secondary structures from the Das lab deposited on their GitHub site: https://github.com/DasLab/FARFAR2-SARS-CoV-2
And for a quick and comprehensive reference of RNA structure, I'd look at this Biopolymers review by Prof. Turner (DOI: 10.1002/bip.22294) and the famous reference "Principles of Nucleic Acid Structure" by Wolfram Saenger, 1984, New York: Springer-Verlag.





#RNA to create the protein.