Summary
Reference materials (RMs) consisting of whole microbial cells characterized for total cell number and total DNA are needed to evaluate and compare cell counting methods, to measure and optimize DNA extraction efficiencies and next generation sequencing (NGS) protocols, and to serve as benchmarks for the probiotic and live microbial therapeutics industries. However, quantifying microbial cells is challenging since microbes vary in size (often < 1 µm in diameter), shape (rod, sphere, etc.) and morphology (single cells, doublets, chains, clusters, etc.), and typically have more than one genome copy per cell. We are establishing measurement methods to quantify total microbial cells and total DNA in a sample. These methods will be used to assign values to future NIST RMs consisting of whole microbial cells. These RMs will enable stakeholders to assess accuracy, bias, and comparability among various quantification approaches.
Description
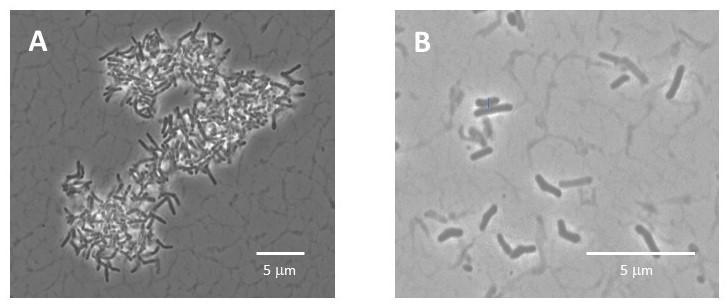
Phase contrast images of Actinomyces naeslundii, a highly aggregated oral bacteria, taken before (A) and after (B) disaggregation using focused ultrasonication.
Bacterial disaggregation as a strategy to improve cell counting (S. Servetas, S. Da Silva)
Counting bacterial cells is challenging enough when they are single, individual cells. It can be further complicated by the various bacterial morphologies (e.g., chains, clusters, etc.), which interfere with accurate bacterial cell enumeration.
We have optimized an approach using focused ultrasonication to disaggregate bacteria chains and clusters with limited damage to the cells as assessed by membrane integrity staining and culture-based methods. Disaggregation has been demonstrated using the highly aggregated Actinomyces naeslundii and the chain-former Streptococcus mitis. Our results indicate focused ultrasonication is a promising technique to disperse bacterial aggregates for improved counting accuracy.
Increasing Confidence in Total Cell Count (J. Dunkers, S. Da Silva, S. Servetas)
Confidence in total cell count requires robust protocols, well-characterized calibrants, and comparison of results with those from orthogonal techniques. We are developing and comparing methods to count microbes using flow cytometry, impedance-based counters (Coulter technology), optical microscopy, and fluorescence plate readers.
Our approach involves first establishing the instrument response for each technique using fluorescent 1 µm diameter calibration beads. The results provide insight into the proportionality of the response as well as accuracy, precision, and upper and lower limits of detection, revealing inherent differences among the counting techniques. Then, the workflow, with the addition of culture-based methods for comparison to the typical colony forming unit (CFU) measurement, is repeated on microbial cells. These efforts will pave the way for high confidence methods to quantify total cell number in pure microbial cell reference materials.
Methods to Measure Total DNA
Lysing bacteria to release DNA is an inefficient process that varies among bacterial strains and background matrix conditions. An independent method to quantify total DNA would provide a much needed baseline to assess and optimize DNA extraction protocols in support of general cell quantification and characterization of extraction biases in metagenomic workflows.
We are evaluating two complementary methods to quantify total DNA. The first approach uses acid/base hydrolysis to release the nucleobases from the DNA and liquid chromatography-mass spectrometry (LC-MS) to quantify the nucleobases in a microbial sample (S. Servetas, S. Forry, M. Lowenthal). The second approach applies flow cytometry and fluorescent DNA-binding dyes to quantify the amount of DNA per cell (or small cell cluster) on a cell by cell basis (S. Da Silva). Results can be coupled with a total cell count to calculate the total amount of DNA in a cell. These methods will help establish the fundamental metrology for quantifying total DNA in a microbial sample.

