Illarion Turko (Fed)
Research Chemist
The scientific expertise of Dr. Turko is in protein chemistry and mass spectrometry. He has authored or co-authored more than 80 peer-reviewed publications. His current research focuses on two major projects: 1) Internal standards for targeted proteomics and 2) Assessing morphology of monoclonal antibody (mAb) aggregation.
(1) Internal Standards for Targeted Proteomics
Targeted proteomics is a technique that has received much attention in academia and in pharmaceutical/biotechnological industries. The greatest benefit of this technique lies in the highly sensitive and quantitative detection of proteins, peptides, and post-translational modifications. Quantification by targeted proteomics relies on mass spectrometry and internal standards, which can be stable isotope-labeled or non-labeled. In addition to traditional standards comprised of either recombinant proteins or synthetic peptides, artificial proteins composed of concatenated tryptic peptides (QconCATs) have recently been introduced as a conceptually new material for use as an internal standard for targeted proteomics. We focus on design, expression, purification, characterization, and application of QconCATs and on side-by-side comparison of QconCATs with full-length proteins and synthetic peptides. Immediate trends in our research include: (i) incorporation of natural flanking sequences for every Q-peptide to make quantification based on QconCATs more accurate (ii) genetic insertion of post-translational modifications (PTMs) into the QconCATs for unlimited quantification of various PTMs. Overall, the use of QconCATs will find applications in quantification of the protein differences that are associated with normal and diseased states, which will ultimately improve our basic knowledge of diseases and will provide input in identification of new therapeutic targets.
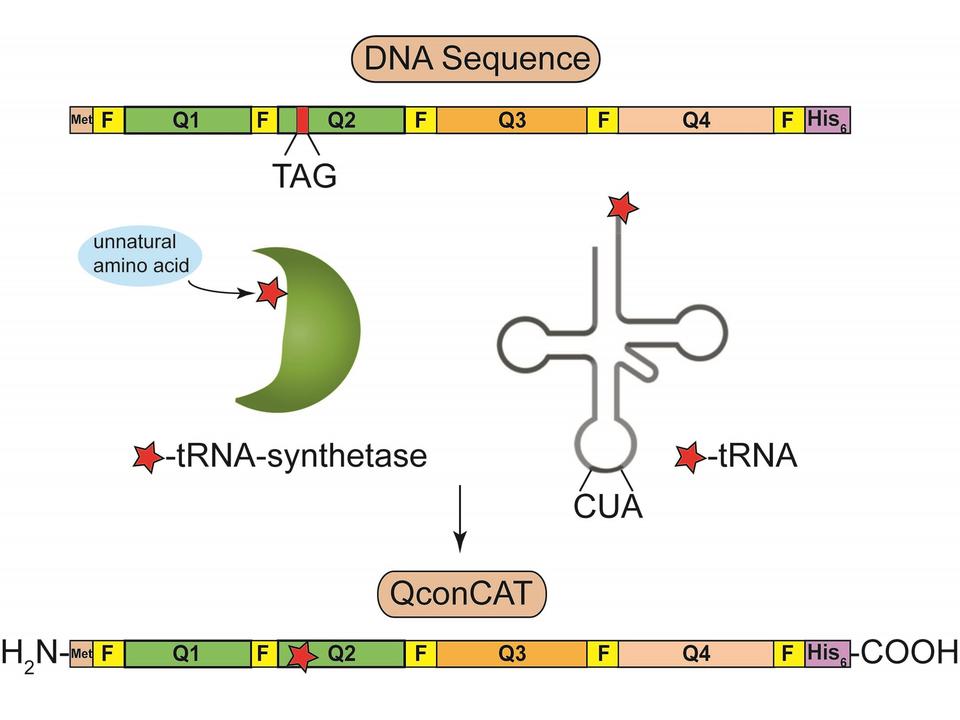
(2) Assessing Morphology of Monoclonal Antibody (mAb) Aggregation
There are many environmental factors that can lead to aggregation of mAbs and the final state of aggregation seems to depend on the aggregation pathway and appears in a variety of forms. The extent to which different forms of mAb aggregates impact biological activity and the risk of immunogenicity is poorly understood, primarily because of the limitations of existing techniques. These techniques assess the size and number of aggregates rather than aggregate morphology. We reasoned that the protein-protein interfaces associated with the mAbs aggregation could be selectively recognized by short peptides with random sequence. In other words, those protein-protein interfaces that do not exist in monomeric mAbs, but do present in aggregated mAbs can be a target for high affinity peptide reagents. This idea directed our attention to peptide phage display technology, which is a powerful tool in the identification of ligands with novel functions. Peptides bound to aggregate interfaces can be selected from a complex mixture of billions of displayed peptides on phage and further enriched through the bio-panning process. Once identified, the selected peptides can be used for developing quantitative methods to assess the morphology of mAb aggregation.

