Programs and Activities
Metabolomics Quality Assurance and Quality Control Materials (MetQual) Program
The NIST Metabolomics Quality Assurance and Quality Control Materials (MetQual) Program has been created to demonstrate and improve the comparability of metabolomics measurements for industry, government, and academic laboratories through the use of NIST Reference Materials and Data (RMDs). The developed RMDs will serve as quality assurance and quality control (QA/QC) materials for routine metabolomic measurements, differential studies, interlaboratory comparisons, laboratory and instrument qualification, and training for the metabolomics community. Furthermore, these RMDs will be assessed by both NIST and community-based interlaboratory comparisons. Contact: christina.jones [at] nist.gov (christina[dot]jones[at]nist[dot]gov)
Gut Microbiome Metabolomics Interlaboratory Program
The Gut Microbiome Metabolomics Interlaboratory Program is an effort within the NIST Microbiome Program. This metabolomics interlaboratory exercise is being developed to help obtain consensus characterization of human whole stool materials and assess measurement variability within the metabolomics community. This program will also inform NIST about the measurement capabilities within the gut microbiome metabolomics communities and assist in the design and characterization of future whole stool reference materials. Contact: paulina.piotrowski [at] nist.gov (paulina[dot]piotrowski[at]nist[dot]gov)
Method Assessment for Non-Targeted Analyses (MANTA) Program
The MANTA Program is an ongoing collaboration with laboratories to increase the comparability and reproducibility of their results using non-targeted analysis techniques, with a specific focus on instrumental techniques and subsequent data analysis. Collaborators include laboratories from academia, government and industry that currently apply non-targeted analytical tools, or are interested in establishing a non-targeted analysis program. The program includes reference material creation, interlaboratory studies, and novel computational tool development. Initially, the program will focus on the use of liquid chromatography with mass spectrometry for non-targeted analysis, with the intent of expanding the program to other techniques. Contact: benjamin.place [at] nist.gov (benjamin[dot]place[at]nist[dot]gov)
Environmental Metabolomics Program
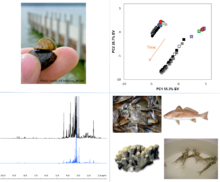
NIST has a well-established NMR-based metabolomics program centered on the unique advantage of a multi-institutional partnership at the Hollings Marine Laboratory (HML) coupled with a high-field nuclear magnetic resonance (NMR) facility. This program leverages the skills and resources of HML researchers with expertise and experience on the biology of a wide variety of aquatic organisms from microbes to marine mammals. By applying NMR, an unbiased discovery tool, to investigate non-model organisms, we can bring new understanding to their biochemistry, while answering focused questions on the impacts of environmental stressors ranging from physical changes in the environment to challenges from industrial and urban pollution. The application of metabolomics to environmental systems biology is broad and diverse, comprising large scale biomonitoring efforts, enhancing aquaculture science, and understanding mechanisms involved in pathophysiology of disease, to name a few. Each biological system bears distinctive challenges for high quality metabolomics research. The knowledge gained from extensive study design and method development in environmental metabolomics research presents a novel opportunity to influence and contribute to best practices, new techniques and protocols and data tools that can be transferred to application in human health. Contact:tracey.johnston [at] nist.gov (tracey[dot]schock[at]nist[dot]gov)
Comparative Mammalian Proteome Aggregator Resource (CoMPARe) Program

NIST is developing a portal to enable researchers to mine high-quality proteomic data from phylogenetically diverse species, to identify advantageous biological adaptations and drive human biomedical breakthroughs. Initially the CoMPARe Program has generated proteomic data from sera from over 30 different species that currently have genome annotations. The resulting data will be publicly available and a data tool has been developed to humanize protein identifications between species to facilitate direct comparisons. In addition to acquiring data on more species, current efforts are focused on a web portal to allow for easy species-species comparisons directed toward expert and non-expert end-users. This will create the foundation for the next phase that will generate data on 50 additional mammalian species as genome annotations become available. This project will be the template for future comparative proteomic projects evaluating plasma, serum and specific tissues of interest in non-model organisms. Contact: benjamin.neely [at] nist.gov (benjamin[dot]neely[at]nist[dot]gov)
Chemical Measurements for Microbial Ecology

Multifaceted microbiome measurements are necessary to identify the molecular mechanism(s) that are responsible for the observed clinical efficacy of probiotics, Fecal Microbiome Transplants (FMTs) and other Live Biotherapeutic Products (LBPs). NIST is supporting the biomanufacturing community by providing materials and reference methodologies for mass spectrometry and NMR based metabolomic measurements of microbiomes to understand the complex biochemical signaling that occurs within the microbial community. Because of the complexity of microbial communities, we are working to develop measurements that will enable more standardized and reproducible microbiome measurements. We are performing studies that evaluate multiple species individually and in a community setting by measuring special interactions, metabolic activity, and fitness as a function of growth conditions. Contact: paulina.piotrowski [at] nist.gov (paulina[dot]piotrowski[at]nist[dot]gov)
Best Practices in Microbial Metabolomics Research
NIST studies fundamentals in the metabolomics workflow to promote best practices for the community. For microbial research, a systematic study addressed impacts of storage solution on metabolomics analysis by NMR. Multiomic studies concerning microbial samples require specific attention to sample preservation solutions when intended for metabolomic analyses. Solutions routinely employed in genomic and transcriptomic studies for nucleic acid stabilization may obscure or alter metabolomic results, or the solutions may be incompatible with platforms like mass spectrometry causing system contamination. An evaluation of sample storage solution impacts on metabolomic profiles outline important considerations for researchers on sampling practices for microbial multiomics studies that incorporate metabolomic analyses. Contact: kehau.usui [at] nist.gov (kehau[dot]usui[at]nist[dot]gov)
Steroid Hormone Pathway Mapping
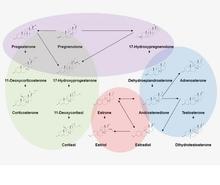
The endocrine system orchestrates major developmental, reproductive, and other physiological changes throughout life. Many techniques use single analyte analysis for clinical diagnosis of endocrine disease, but steroids are produced in cascades that can elucidate action pathways. Mapping of steroid hormone pathways can improve understanding and diagnosis of endocrine diseases and endocrine disruption. For more information, follow the links on steroid pathway mapping, human standard reference materials related to clinical hormone measurements, and application to environmental and agricultural endocrinology. Contact: ashley.boggs [at] nist.gov (ashley[dot]boggs[at]nist[dot]gov)

