Structure of CMV Genome
Structure of Cytomegalovirus Genome
Simplified genomic structure:

Schematic map of the CMV genome. The CMV genome is organized as two regions of unique sequences, unique long (UL) and unique short (US), flanked by two sets of inverted repeats (TRL/IRL) and (IRS/TRS) (light shaded boxes). Kotenko et al. PNAS February 15, 2000 vol 97 no. 4 pp. 1695-1700
Nomenclature:
- UL – Unique long
- US – Unique short
- TRL – Terminal repeat long
- IRL – Internal repeat long; inverted repeat of TRL
- TRS –Terminal repeat short
- IRS – Internal repeat short; inverted repeat of TRS
Detailed genomic structure:

Map of the HCMV genome. The green arrows represent previously annotated ORFs of HCMV likely to encode a bona fide polypeptide, red arrows designate ORFs unlikely to encode a polypeptide, and blue arrows indicated previously unrecognized ORFs that the present analysis predicts have high potential to encode proteins. The gray box marks the additional sequence found in the HCMV Toledo strain, locating it with respect to the AD169 genome. Rectangles superimposed on the line represent the sequence-identify terminal repeats. Each mark on the sequence line represents 1,000 bp. Murphy et al. PNAS November 11, 2003 vol 100 no 23.
Alternate structures
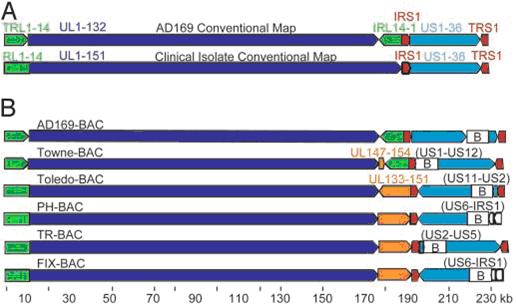
HCMV ORF organization.
(A) Conventional ORF maps of the AD169 laboratory strain and clinical isolates. From the left, the AD169 genome contains TRL1-14 (green arrow), UL1-132 (dark blue arrow), IRL14-1 (green arrow), IRS1 (red arrow), US1-36 (light blue arrow), and TRS1 (red arrow). From the left, clinical isolates contain a unique domain including RL1-14 and UL1-151 (green plus dark blue arrow) followed by IRS1 (red arrow), US1-36 (light blue arrow), and TRS1 (red arrow).
(B) ORF maps of the BAC clones whose sequences are reported in this article (see citation below). Sequences were linearlized at the position corresponding to nucleotide 1 of the original AD169 sequence. Arrows indicate the relative orientations of the repeated and unique ORF blocks. The RL region is not repeated in the clinical isolates (Toledo, PH, TR, and FIX); there is a single copy of the region (green segment appended to the unique long region). The Towne laboratory strain contains a block of ORFs (UL147-154, orange arrow) that is not present in AD169, and the clinical isolates contain a block of ORFs (UL133-151, orange arrow) that is not present in the AD169 laboratory strains. The BAC inserts are identified (B), and viral ORFs deleted during BAC insertions are listed in parentheses. Murphy et al. PNAS December 9, 2003 vol 100 no 25.
Contacts
Applied Genetics Group
-
(301) 975-5473

