Summary
Metagenomics, or identifying the organisms present in a sample through DNA-based analysis, is a revolutionary technology made possible through advances in sequencing technologies and computing power. Like all sample analyses, standards are needed to benchmark performance and enable translation into applications such as clinical diagnostics, microbiome therapeutics, environment, and bio-surveillance. Given the complexities of the analyses, an entire suite of NIST Reference Materials are under development to provide measurement assurance.
Description
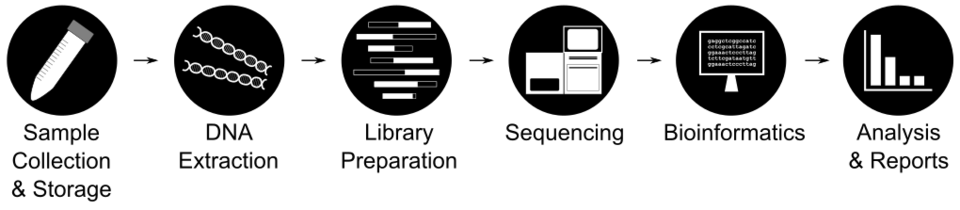
Metagenomics involve a series of steps, including sample collection, extraction, DNA preparation, sequencing, bioinformatics, and reporting. Bias at each step contributes to error in the final analysis. With a wide variety of options for each step, two analyses can produce different results. Standards can help characterize the biases, enabling developers and regulators to recognize any limitations and have confidence in the results.
Metagenomics is a powerful technology combining high-throughput (a.k.a. “next-generation”) sequencing with advanced bioinformatics to identify and quantify mixtures of organisms in a sample. Metagenomics workflows consist of 4 main steps:
- Sample collection: location, amount, storage, etc.
- Extraction: method, amount
- Sequencing: preparation, platform, chemistry
- Bioinformatics Analysis: algorithm, database, pre- and post-filtering
Each step contributes some error, or bias, to the overall measurement, which in some cases can result in wildly different results from what is known or expected. Error propagation is linear, which offers an opportunity to characterize it by working backwards using well-characterized materials suited for benchmarking each critical step.
DNA-based Reference Materials
Two DNA-based materials (RM 8375, a 4-bacteria panel, and RM 8376, a 19 bacteria + 1 human panel) were designed around sequencing and analysis, and are known abundance (chromosomal copy number concentration) with high-quality draft genome assemblies. These enable side-by-side in silico and wet-lab characterization of the metagenomic analysis, and can serving as ground truth to assist validation of portions of the metagenomic workflow.
These and other nucleic acid standards are available for purchase through the Office of Reference Materials website.
Whole -Cell Reference Materials
Whole cell materials to help characterize extraction efficiencies are also under development. 5 individual organisms and 1 organism mixture sample sets will each be homogeneous and stable with respect to cell number. The 5 organisms represent Gram-positive and negative strains and will enable intercomparability between laboratory protocols.

