Summary
Measurements of the performance of engineered organisms are typically not reproducible or comparable across different laboratories. So, to enable meaningful comparison, exchange, and operation of engineered organisms across organizations, we are working to create a framework for traceable, comparable measurements of transcription rates in engineered microbial systems.
Description
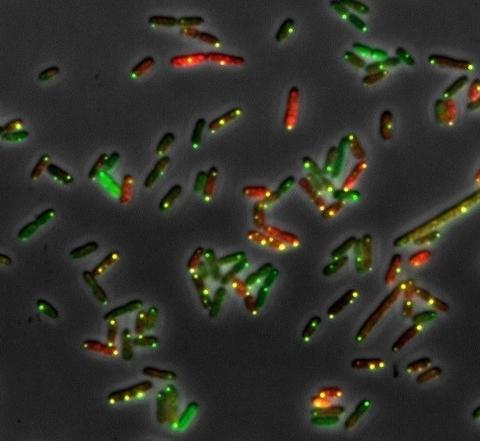
Multi-color fluorescence microscopy for absolute quantitation of mRNA molecules in engineered bacterial cells. Image credit: Jayan Rammohan and Bin Shao, MIT/NIST.
The ability to engineer novel, useful functions into microbial cells for manufacturing or therapeutic applications has accelerated rapidly over the last few decades. However, the measurements required to underpin predictive engineering of these systems are typically not comparable across different laboratories. Consequently, the engineering of biology remains a laborious and costly trial-and-error based process.
To enable data sharing for predictive design and modeling of engineered organisms, and to provide quantitative metrics for genetic circuit performance, NIST is developing methods to count individual mRNA molecules in engineered bacterial cells. These methods are based on a technique known as single molecule RNA FISH. Counting of mRNA molecules in single cells has several advantages over conventional population-average based mRNA measurements, including traceability to the International System of Units, and the ability to determine the distribution of transcription levels across a population of cells. NIST is also working on comparisons of the different variants of the RNA-FISH method, and automation of the RNA-FISH sample preparation process.
RELATED PUBLICATIONS
Shao B, Rammohan J, Anderson DA, Alperovich N, Ross D, Voigt CA. Single-cell measurement of plasmid copy number and promoter activity. Nat Commun. 2021 Mar 5;12(1):1475. doi: 10.1038/s41467-021-21734-y. PMID: 33674569; PMCID: PMC7935883.
Rammohan J, Lund SP, Alperovich N, Paralanov V, Strychalski EA, Ross D. Comparison of bias and resolvability in single-cell and single-transcript methods. Commun Biol. 2021 Jun 2;4(1):659. doi: 10.1038/s42003-021-02138-6. PMID: 34079048; PMCID: PMC8172639.

