Development of NIST Reference Materials for Untargeted Metabolomics, Lipidomics and Proteomics
Summary
The use of metabolite, lipid, and protein profiles for systems biology, biomedical research, and clinical applications has risen dramatically over recent years. Global profiling allows for the detection, identification, and relative quantification of candidate biomarkers and molecular markers. These compounds discriminate phenotypic functions in disease processes, oxidative stress, injury, histopathological changes, pathophysiological mechanisms, drug effects, and glycosylation. It is now possible to observe 1000s to 10,000s of unknown chemical compounds and different protein groups from cell lysates, bio-fluids, and tissue samples. This untargeted data analysis approach can highlight gross differences between sample groups and can be further queried by targeted quantitative analysis for specific proteins and/or biomarkers of interest. It is challenging, however, to compare data across instrument runs or between studies without a common internal standard or a matrix-matched QC check, and NIST is working to provide relevant reference materials to address this need.
Description
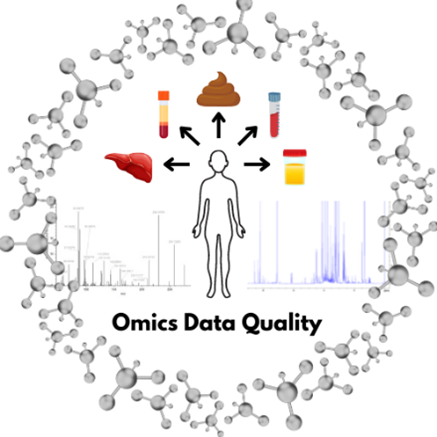

There is a significant need for tools to increase harmonization, comparability, and reproducibility between laboratories and/or studies, and to provide quality control of sample preparation and data analysis protocols utilized in untargeted omics analysis. Reference materials are being developed to standardize quality assurance/quality control (QA/QC) materials and support the development of methods for characterizing new technologies. In addition, these reference materials will facilitate the generation of publicly available reference datasets, which can be used to evaluate data processing and analysis pipelines.
Current Activity
NIST is developing standalone RMs and RM suites including plasma, urine, and liver matrices representing different phenotypes and health states. NIST is also generating system suitability extracts from bio-fluid and tissue-based materials to develop best practices and to facilitate harmonization of NMR- and MS-based metabolomic and lipidomic measurements. Datasets are being created with highly confident identifications of chemical compounds, peptides, and protein groups. These datasets will be used to validate analytical data processing and results for untargeted studies.
STANDALONE REFERENCE MATERIALS
NIST SRM 1950 – Metabolites in Frozen Human Plasma: SRM 1950 was the first reference material developed specifically for metabolomics research. This SRM represents ‘normal’ human plasma and was obtained from 100 individuals. Plasma from an equal number of men and women in a narrow adult age range (40 to 50 years) was used. Racial distribution of the donors reflected the distribution in the U.S. population. Contact: johanna.camara [at] nist.gov (johanna[dot]camara[at]nist[dot]gov)
NIST RM 8461 – Human Liver for Proteomics: RM 8461 was produced to provide a complex human tissue sample for evaluating the performance of proteomics mass spectrometry instruments coupled to liquid chromatography (LC), software for the identification of proteins and peptides, in addition to quality control in omics studies. Contact: clay.davis [at] nist.gov (clay[dot]davis[at]nist[dot]gov) or benjamin.neely [at] nist.gov (benjamin[dot]neely[at]nist[dot]gov)
reference material suites
NIST Candidate RM 8231 – Frozen Human Plasma Suite for Metabolomics: The new suite will consist of pooled plasma from diabetic, high triglyceride, and young African American plasma samples. RM 8231 provides a range of constituent metabolite and protein profiles. Contact: tracey.schock [at] nist.gov (tracey[dot]schock[at]nist[dot]gov) and clay.davis [at] nist.gov (clay[dot]davis[at]nist[dot]gov)
NIST Candidate RM 8232 – Frozen Human Urine Suite for Metabolomics: The suite will consist of female non-smoker, female, smoker, male non-smoker, and male smoker pooled samples. The suite will be qualitatively characterized using liquid chromatography–high resolution mass spectrometry and NMR. Contact: tracey.schock [at] nist.gov (tracey[dot]schock[at]nist[dot]gov) and clay.davis [at] nist.gov (clay[dot]davis[at]nist[dot]gov)
NIST Candidate RM 8462 - Human Liver Suite for Proteomics and Metabolomics: RM 8462 is being designed to maximize expected variability in the observed proteome and metabolome. The material suite will consist of normal, fatty, and congested liver samples and both protein and metabolite fold changes and percent differences for the top proteins and metabolites contributing to differentiation will be determined. Contact: clay.davis [at] nist.gov (clay[dot]davis[at]nist[dot]gov)
NIST Candidate RM 8048 - Human Whole Stool Reference Material: RM 8048 is being developed to enable comparability and improve reproducibility in metagenomics and metabolomics analysis. Contact: scott.jackson [at] nist.gov (scott[dot]jackson[at]nist[dot]gov) For more information also see the Human Gut Microbiome Reference Material.

