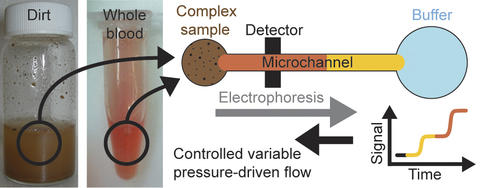
Illustration of a GEMBE device: a complex biological sample such as dissolved dirt or whole blood is pushed by an electric field toward a microchannel. Buffer fluid flowing in the opposite direction acts as a gate. Gradually reducing the buffer flow slowly 'opens' the gate, allowing individual components from the sample to enter the microchannel when the pressure becomes weaker than the electric force pushing each component molecule. These molecules then travel past a detector that analyzes them. Unwanted components are kept out of the microchannel.
It's not often that someone can claim that going from a positive to a negative is a step forward, but that's the case for a team of scientists from the National Institute of Standards and Technology (NIST) and private industry. In a recent paper, the group significantly extended the reach of their novel microfluidic system for analyzing the chemical components of complex samples. The new work shows how the system, meant to analyze real-world, crude mixtures such as dirt or whole blood, can work for negatively charged components as well as it has in the past for positively charged ones.
In previous work, NIST researchers Elizabeth Strychalski and David Ross, in collaboration with Alyssa Henry of Applied Research Associates Inc. (Alexandria, Va.), demonstrated the use of a technique called GEMBE (for "gradient elution moving boundary electrophoresis") for analyzing complex samples. The NIST-developed system combines a simple microfluidic structure (two reservoirs connected by a microchannel), electrophoresis (which uses electricity to move sample components through a fluid) and pressure-driven flow. (See "'No Muss, No Fuss' Miniaturized Analysis for Complex Samples Developed.")
Analyzing complex samples can be difficult because components in these samples (such as the fat globules in milk or proteins in blood) can "foul" or contaminate microfluidic channels. The traditional solution has been to remove contaminants with costly, time-consuming sample preparation prior to analysis.
GEMBE solves this problem by pumping fluid through the microchannel using a controlled pressure in the direction opposite to electrophoresis. This opposing pressure-driven flow acts as a "fluid gate" between the sample reservoir and the microchannel. Gradually reducing the pressure of the counterflow opens the "gate" a little bit at a time. A specific sample component is detected when the pressure flow becomes weak enough—i.e. the "gate" opens wide enough—that the component's electrophoretic motion pushes it against the pressure-driven flow and into the channel for detection. In this way, different components enter the channel at different times, based on their particular electrophoretic motion. Most importantly, the channel doesn't become fouled because the unwanted components in the sample are held out.
"Previously, we validated the GEMBE technique by quantitatively analyzing components from complex samples in solution that were cationic [positively charged] and could, therefore, be separated relatively easily from anionic [negatively charged] contaminants in a mixture," Strychalski says. "However, we needed a way to make GEMBE work when both the desired components and the contaminants are negatively charged."
For some samples, Strychalski says, this was achieved by choosing a different solution pH to change the electrophoretic motion of the unwanted components. In other cases, the addition of commercially available surface coatings to the sample did the trick without compromising the ease and robustness of the GEMBE technique.
"Additives can be selected that will interact with material in the sample that we don't want to study," Strychalski explains. "If we choose the right coating, it will slow the electrophoretic motion of contaminants relative to the desired components. This prevents the former from interfering with analysis while still allowing the latter to enter the microchannel for detection."
Strychalski and her colleagues plan to continue refining the GEMBE system, including an effort to define which surface coatings optimize the technique for specific components in a variety of complex samples.

