NIST's Integrated Colony Enumerator (NICE)
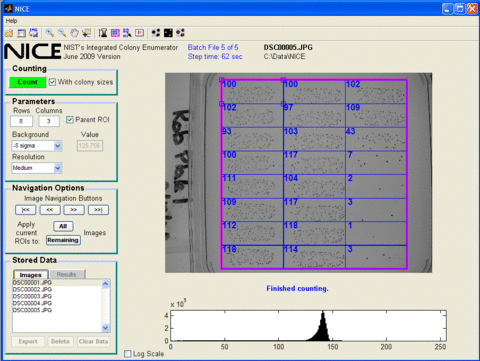
Enumeration of bacterial colonies on an agar plate is simple in concept, but automated colony counting is difficult due to variations in colony color, size, shape, contrast, and density, as well as colony overlap. Furthermore, in applications where high throughput is essential, it is critical to employ a fast and user-friendly automated technique that does not compromise counting accuracy. We have developed a colony counting program, NIST's Integrated Colony Enumerator (NICE), designed to count dark colonies from multiple regions of interests on an agar plate. Images can be cost effectively acquired by a digital camera or a desktop scanner and imported into NICE. Multiplexed, high throughput standardized assay formats such as the multiplexed opsonophagocytic killing (MOPA4) assay can be readily counted by NICE.
There are two software files required to run NICE. There is also one supplementary file and documentation. Descriptions of these files are available below, and the files can be downloaded from the following ftp site, click here to download.
Related Publications
Contact
-
(303) 497-7287
-
(303) 497-6588

