Summary
Characterizing and predicting cell population behavior is critical to the efficiency and flexibility of manufacturing processes of cell-based products. Characterization is challenging because individual cells within populations demonstrate heterogeneous and dynamic characteristics, and it is often unclear what measurements provide meaningful and predictive information about important population indicators of critical activities such as proliferative capacity or differentiation potential.
Description
Heterogeneity of cell populations indicates the many ways cells can process information.
We study heterogeneity in gene expression by inserting a gene for a fluorescent protein downstream of the gene we are interested in, and we examine large numbers of cells by imaging or flow cytometry. We think of the heterogeneous distributions of responses in a cell population (Panel A) as analogous to a potential energy landscape (Panel B).
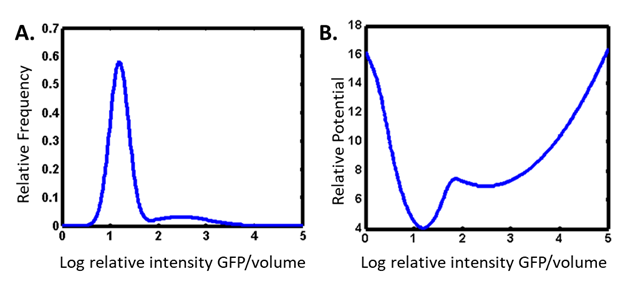
This theoretical construct allows us to model biological networks as nonequilibrium thermodynamic systems. This approach provides a window into which are the most important characteristic measurands of the population. Quantitative imaging of individual cells over time allows us to determine the rate with which cells fluctuate in their expression of the probe.
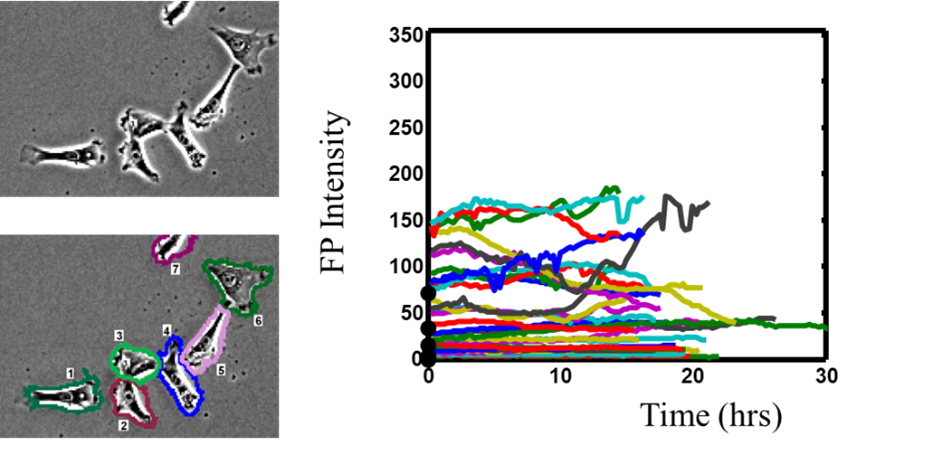
We then use that dynamic information in a stochastic model to predict the rates of response of the population to perturbations (1).
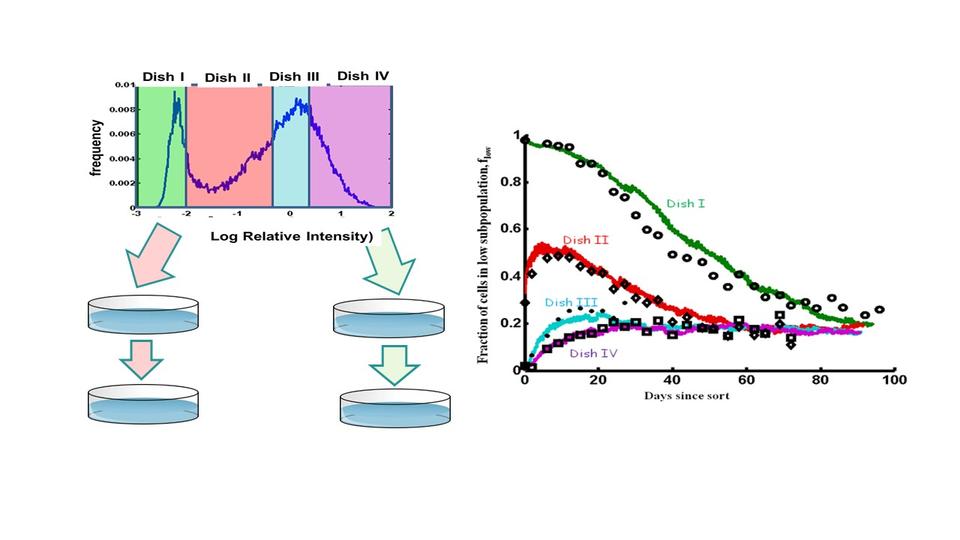
Statistical thermodynamics modeling can provide a path for identifying a small number of the most important cellular features that can be easily measured on the manufacturing floor. Developing these theoretic models requires the development of advanced methods to enable the collection of very large datasets where many cells are sampled at relatively high frequencies.
Advancing technologies in imaging, image analysis and theoretical modeling
Measuring the temporal expression of more than one fluorescent reporter simultaneously in individual cells can provide data that indicate strong correlations – causative relationships – between network components (2).
The relationships between transcription factors that control pluripotency and differentiation will likely produce a 2-dimensional landscape like this:
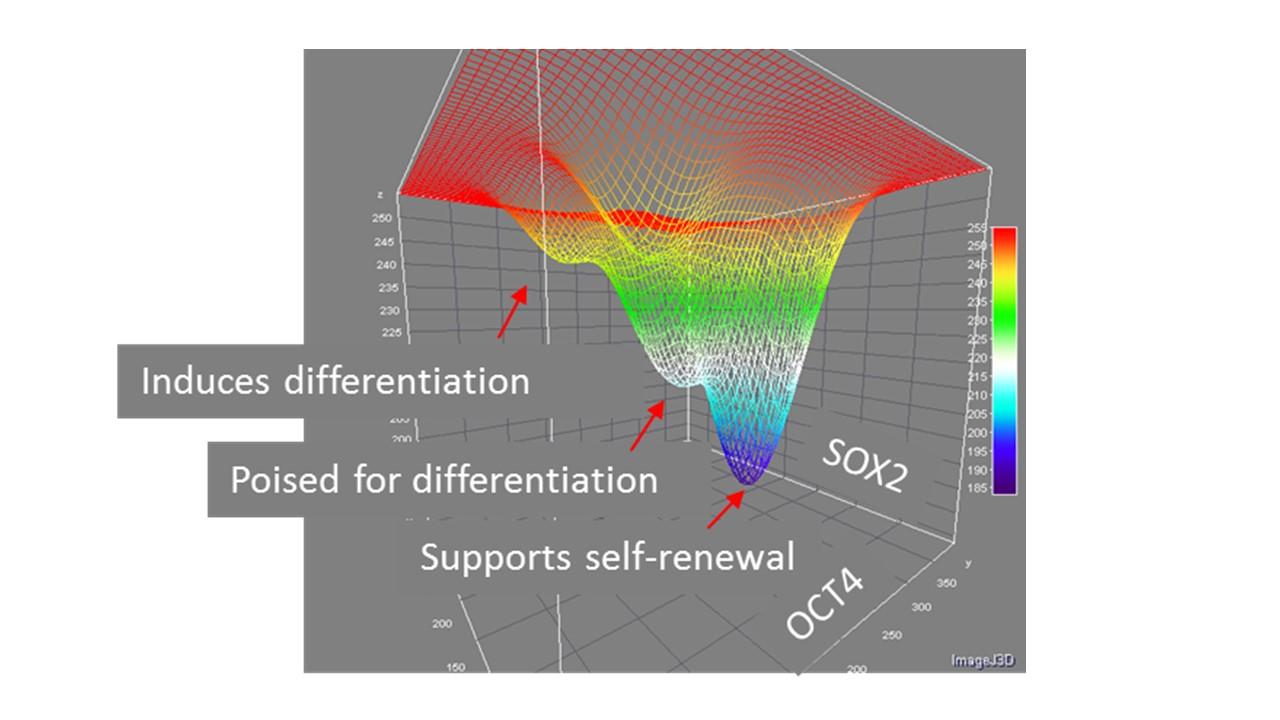
Quantifying temporal fluctuations of reporter molecules in Induced pluripotent stem cells requires segmenting cells that exist very close to one another in colonies and tracking them over long times as they divide. This requires imaging each cell at a rate of about every 2 minutes, making it necessary to minimize the use of fluorescence and to train AI models to recognize and track cells in bright field imaging. And of course, one needs to be able to sample thousands of cells within that time frame, resulting in very large volumes of data Advances in imaging and image analysis capabilities are under development in support of this data-driven theoretical approach.
CITATIONS
- Sisan D.R., Halter M., Hubbard J.B., Plant A.L. (2012) Predicting rates of cell state change due to stochastic fluctuations using a data-driven landscape model. PNAS 109, 19262-19267. https://doi.org/10.1073/pnas.1207544109
- Hubbard, J.B, Halter, M., Sarkar, S., Plant, A.L. (2020) The Role of Fluctuations in Determining Cellular Network Thermodynamics. PLoS. https://doi.org/10.1371/journal.pone.0230076

