Description
The science of human health and disease often includes studies of cells, the smallest unit of biological complexity. Microscopic imaging is an important tool in these studies and can provide morphological and molecular information. Molecular microscopic data is most often collected in a static mode after external addition of fluorescent labels that bind to cellular molecules of interest. A more powerful, but more technically difficult approach is to image living cells over time in order to observe dynamic changes in morphology, molecular concentration and location, and stage of cell division (1, 2). Molecular information in this case is generally provided by the genetic engineering of cell lines that express fluorescent proteins within the living cells.
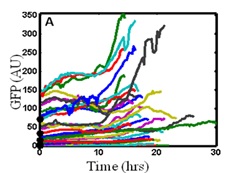
We are developing statistical thermodynamics models that are based on dynamic fluctuations in the activity of select genes which are reported on by fluorescent proteins. We have performed such an analysis in the past (3), and the results of the analysis allowed accurate prediction of cell population responses. Shown on the left is a sample of fluorescent trajectories for individual cells, from which mean square displacements were calculated, providing an average diffusion coefficient for fluctuations in the expression of that gene. We have recently developed a similar theoretical approach for multivariable gene systems (4). We envision (5) a metrology system in which engineered cell lines, advanced imaging techniques, and a stochastic modeling framework will allow the characterization, monitoring, and prediction of cell populations to aid the development of manufacturing pipelines.
We are currently designing cell lines that will express different fluorescent proteins for multiple genes simultaneously. We are producing pluripotent stem cell lines that express fluorescent proteins in response to their production of 3 of the canonical molecules associated with pluripotency. These cell lines will be used to better understand how these molecules behave in a coordinated fashion to control pluripotency using a theoretical approach based on statistical mechanics. They will also be useful tools to direct manufacturing processes by reporting on how cellular growth conditions influence the stability of the pluripotent state and the tendency of cells to differentiate into specific desired cell types. We anticipate collaborating with others on the use of these cell lines to evaluate cell manufacturing pipelines and providing the cell lines themselves as reference materials.
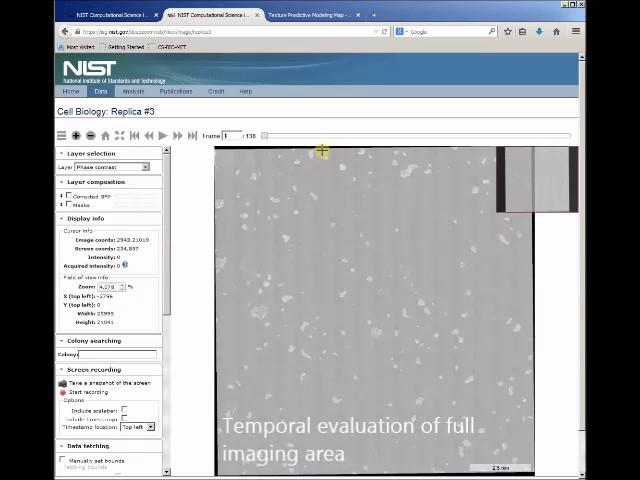
Currently, there are no turn-key imaging systems that make these kinds of measurements easy. To collect and extract high quality data short-time-dependent data from live cell imaging requires rapid scanning for collecting large image datasets and high signal sensitivity. Coupled to the imaging capability is the need for reliable and validated image analysis pipelines. Our programmatic focus is to provide web-based tools (5), instrument benchmarking materials (6), algorithms for segmentation and tracking cells (7-11) and other methods (12), and demonstration of impact in the biomanufacturing of cell-based therapies (13, 14). The figure of stem cell colonies at left, which shows a composite image of many fields of view and is part of a dataset that can be observed at different scales and over time, provides an example of these capabilities applied to stem cells (15). See also https://www.nist.gov/video/wipp-map-app-track-stem-cells
There are additional challenges for imaging living cells to collect meaningful biological information. Biologically relevant conditions (temperature, atmospheric and solution composition) must be maintained. Optical sampling must minimize light-induced damage of cells while collecting image data of sufficient quality with respect to spatial resolution, signal to noise ratio, focus plane, and time resolution. In addition, because every cell can behave slightly differently from other cell, image data sets need to be sufficiently large enough to provide statistical confidence that the population has been adequately sampled. These challenges are often convolved. For example, the composition of the solution the cells are in that required for cell health can be a source of background fluorescence or other optical perturbations. Also, minimizing damage to exposure to incident light may sacrifice signal intensity. The conditions required to achieve biologically relevant imaging conditions will depend on cell type and the type of data of interest.
RELATED PUBLICATIONS
- Halter M, Elliott JT, Hubbard JB, Tona A, Plant AL. Cell volume distributions reveal cell growth rates and division times. JTheorBiol. 2009;257(1):124-30.
- Halter M, Sisan DR, Chalfoun J, Stottrup BL, Cardone A, Dima AA, et al. Cell cycle dependent TN-C promoter activity determined by live cell imaging. Cytometry A. 2011;79A(3):192-202.
- Sisan DR, Halter M, Hubbard JB, Plant AL. Predicting rates of cell state change caused by stochastic fluctuations using a data-driven landscape model. Proc Natl Acad Sci U S A. 2012;109(47):19262-7.
- Hubbard, J.B, Halter, M., Sarkar, S., Plant, A.L. (2020) The Role of Fluctuations in Determining Cellular Network Thermodynamics. PLoS. https://doi.org/10.1371/journal.pone.0230076
- Plant A, Halter M, Stinson J. Probing Pluripotency Gene Regulatory Networks with Quantitative Live Cell Imaging. Computational and Structural Biology Journal. 2020; https://doi.org/10.1016/j.csbj.2020.09.025
- Bajcsy P, Chalfoun J, simon M. Web Microanalysis of Big Image Data. 1 ed: Springer International Publishing; 2018. 197 p.
- Halter M, Bier E, DeRose PC, Cooksey GA, Choquette SJ, Plant AL, et al. An automated protocol for performance benchmarking a widefield fluorescence microscope. Cytometry A. 2014;85(11):978-85.
- Bajcsy P, Cardone A, Chalfoun J, Halter M, Juba D, Kociolek M, et al. Survey statistics of automated segmentations applied to optical imaging of mammalian cells. BMC Bioinformatics. 2015;16:330.
- Chalfoun J, Cardone A, Dima AA, Allen DP, Halter MW. Overlap-Based Cell Tracker. J Res Natl Inst Stand Technol. 2010;115(6):477-86.
- Chalfoun J, Kociolek M, Dima A, Halter M, Cardone A, Peskin A, et al. Segmenting time-lapse phase contrast images of adjacent NIH 3T3 cells. J Microsc. 2013;249(1):41-52.
- Chalfoun J, Majurski M, Dima A, Halter M, Bhadriraju K, Brady M. Lineage mapper: A versatile cell and particle tracker. Sci Rep. 2016;6:36984.
- Dima AA, Elliott JT, Filliben JJ, Halter M, Peskin A, Bernal J, et al. Comparison of segmentation algorithms for fluorescence microscopy images of cells. Cytometry A. 2011;79(7):545-59.
- Plant AL, Elliott JT, Tona A, McDaniel D, Langenbach KJ. Tools for quantitative and validated measurements of cells. Methods Mol Biol. 2007;356:95-107.
- Plant AL, Parker GC. Translating stem cell research from the bench to the clinic: a need for better quality data. Stem Cells Dev. 2013;22(18):2457-8.
- Williams DJ, Archer R, Archibald P, Bantounas I, Baptista R, Barker R, et al. Comparability: manufacturing, characterization and controls, report of a UK Regenerative Medicine Platform Pluripotent Stem Cell Platform Workshop, Trinity Hall, Cambridge, 14-15 September 2015. Regen Med. 2016;11(5):483-92.
- Bhadriraju K, Halter M, Amelot J, Bajcsy P, Chalfoun J, Vandecreme A, et al. Large-scale time-lapse microscopy of Oct4 expression in human embryonic stem cell colonies. Stem Cell Res. 2016;17(1):122-9.

