Summary
Directed Self Assembly (DSA) is one of the leading candidates for next generation lithography for the semiconductor and data storage industries. DSA combines top down and bottom up patterning to provide both sub-10 nm nanostructures and controlled placement. The feature sizes are determined by chemistry through the lengths of the molecules. The placement is determined by a template fabricated by conventional lithography methods. The template epitaxially guides the self assembly of the block copolymer. The spacing of the template is an integer multiple of the natural spacing of the block copolymer. The smaller pitch block copolymer essentially fills in the gaps between the template guides, multiplying the frequency of the conventional pattern. This project focuses on key materials by design issues for developing new block copolymers with even smaller natural spacings that meet the industry criteria of low defectivity, small spacing, etch selectivity, and ease of DSA. We combine molecular simulations with new X-ray scattering methods to validate the simulations on new materials systems and to extend the amount of information that can be extracted from the X-ray measurements.
Description
One of the key issues for DSA is the three dimensional structure of the block copolymer. For the example of line-space patterns made using lamella-forming block copolymers, it is possible to have the DSA form tilted lamella or complex mixed morphologies that are a combination of vertical and horizontal lamella.(1) Many of these morphologies look similar at the top surface where most metrology methods are sensitive.(2-4) The densities of the two blocks are usually similar, resulting in very little contrast between the two blocks in electron and X-ray based measurements such as transmission electron microscopy and X-ray diffraction. The inability to characterize the buried structure of the block copolymer morphology is a critical issue.
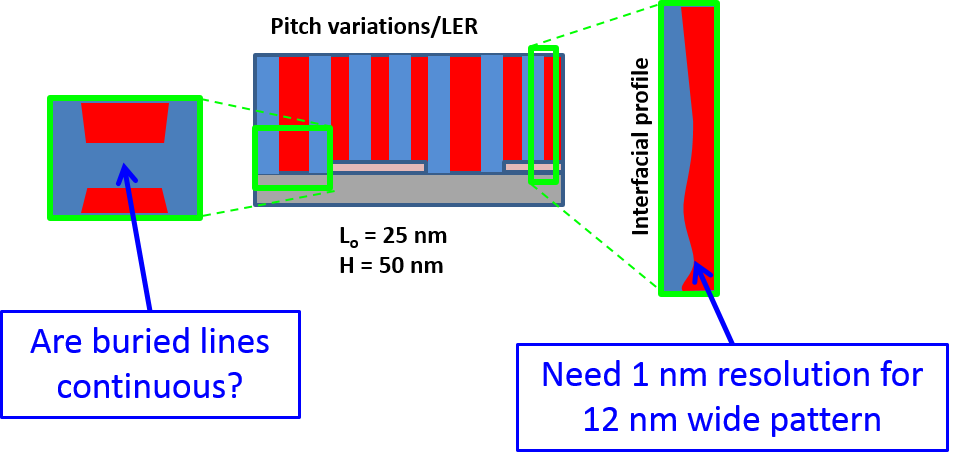
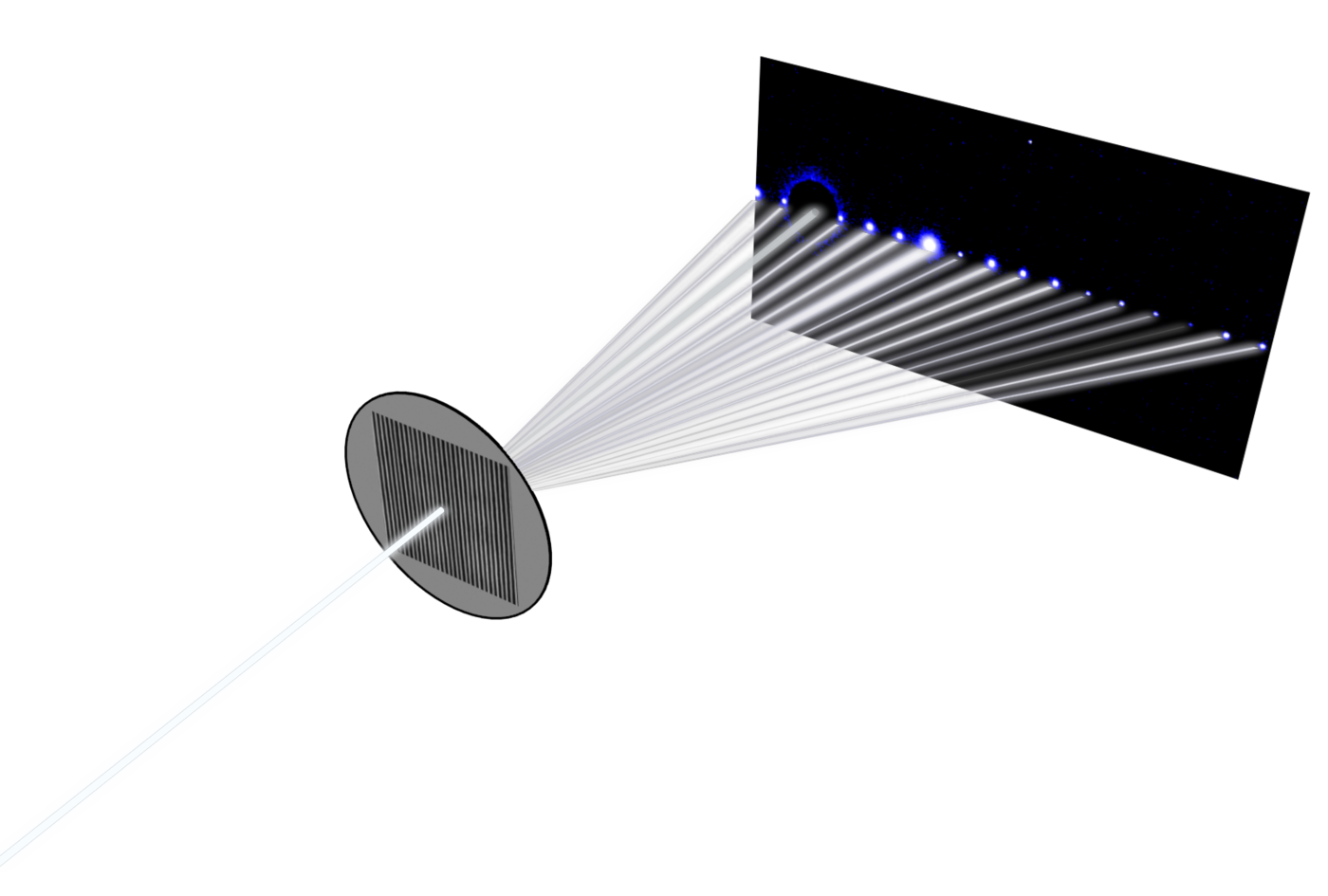
We have developed a new method of X-ray scattering that utilizes resonant soft X-rays with chemical bond sensitivity.(3,4) The method is called resonant critical dimension small angle X-ray scattering (res-CDSAXS) and is based off of the CDSAXS method that we developed for measuring periodic nanostructures.(5) The raw data of the scattering method cannot be directly converted into the original nanostructure and must be modeled using an iterative, inverse method with trial shape solutions. The many parameters required to describe complex shapes make solutions to the inverse problem non-trivial and potentially not unique. This project aims to integrate molecular simulations of the DSA process into the scattering data modeling to constrain the parameters space and maximize the extracted shape information from the scattering measurement. Comparison between the scattering results and molecular simulations also will be used for tuning the simulations for new material systems.
References:
- “Chemical Patterns for Directed Self-Assembly of Lamellae-Forming Block Copolymers with Density Multiplication of Features,” C.C. Liu, A. Ramírez-Hernández, E. Han, G.S.W. Craig, Y. Tada, H. Yoshida, H. Kang, S. Ji, P. Gopalan, J.J. de Pablo, and P.F. Nealey, Macromolecules 46 (4), 2013 1415-1424
- “Graphoepitaxial assembly of cylinder forming block copolymers in cylindrical holes,” B.L. Peters, B. Rathsack, M. Somervell, T. Nakano, G. Schmid, J.J. de Pablo, J. Poly. Phys., Part B, 2014
- “3D X-ray Metrology for Block Copolymer Lithography Line-Space Patterns,” D.F. Sunday, M.R. Hammond, C. Wang, W.L. Wu, G.E. Stein, R.J. Kline, J. Micro/Nanolithography, MEMS, and MOEMS, 12 (3), 2013, 031103
- “Determination of the Internal Morphology of Nanostructures Patterned by Directed Self Assembly,” D.F. Sunday, M.R. Hammond, C. Wang, W.L. Wu, D.M. DeLongchamp, M. Tjio, J.Y. Cheng, J.W. Pitera, R.J. Kline, ACS Nano, 8 (8), 2014, 8426-8437
- “Template-Polymer Commensurability and Directed Self-Assembly Block Copolymer Lithography,” D.F. Sunday, E. Ashley, L. Wan, K.C. Patel, R. Ruiz, R.J. Kline, J. Poly. Phys., Part B, 53 (8), 2015, 595-603

