RFPs: The Best and the Brightest
How, precisely, does disease begin and progress in a single cell? To fully understand such processes on the smallest scales, scientists need a way to peer deep inside the cell and observe how its various components move, change, and interact over time. One powerful tool for that purpose, fluorescence microscopy, can track the dynamics of structures as small as single molecules by "tagging" them with a fluorescent protein (FP)* that emits visible light of a distinctive color when excited by photons from lasers or other sources.
Researchers have used FPs to gain enormous insight into cell biology, but the technique is still limited.
"We can easily see lots of things that occur at high concentrations, in the neighborhood of a million copies, such as the proteins that contribute to cell shape and structure," says Ralph Jimenez of JILA, a joint research center of NIST and the University of Colorado at Boulder. "But if you really want to study the dynamics of living things, it's critical to be able to tag and make movies of cellular components at all concentrations down to single molecules, in a whole organism.
"Right now, it's very difficult to perform time-lapse imaging of cellular constituents present at low concentrations — below, say, 1000 copies," Jimenez continues. "As a result, there's a lot of biology that's invisible to us because we need brighter and more stable red FPs to reveal it. For example, all of the receptors that trigger inflammation and cell death occur at very low concentrations, and we can't examine them at present."
That's why Jimenez's group is using a combination of directed evolution and unique biophysical measurement technology to identify and isolate the next-generation standards for red FPs (RFPs). Red is the group's color of choice because it propagates so readily through living tissue, so it enables observations in tissues where green FPs can't be used.
Several factors affect FP performance. One is photobleaching, which occurs when the molecule has been exposed to so much photon excitation that it finally loses its ability to fluoresce. Another is fluorescence lifetime, or how long the FP will continue to fluoresce before it burns out. The goal is to find RFPs with minimal photobleaching and maximum lifetime (among other properties), which are thus best suited to use in making "molecular movies" of intracellular activity.
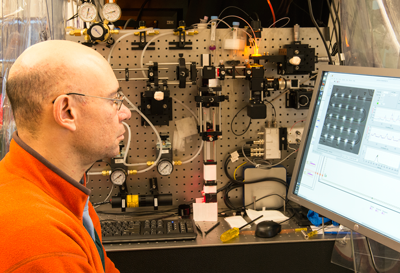

RFPs were first developed about 15 years ago, producing a brief flurry of promising candidates. "They found all the low-hanging fruit," Jimenez says, and "much of the work derived from things that people discovered serendipitously. We're trying to introduce ways to search systematically."
To do that, the team developed new technology with which they can look at very large numbers of RFPs in a short time in very precise spectroscopic detail.
"You'd like to have an RFP that can absorb and emit millions of times before it goes dead, and do it repeatedly and predictably," Jimenez says. "In general, these things are too complicated to design. So the way we approach the problem is to use directed evolution. We take an RFP that more or less behaves, and we modify it based on random mutations or on knowing which regions of the gene are important. Then we can make a million or so different variations on the original and test each of them."
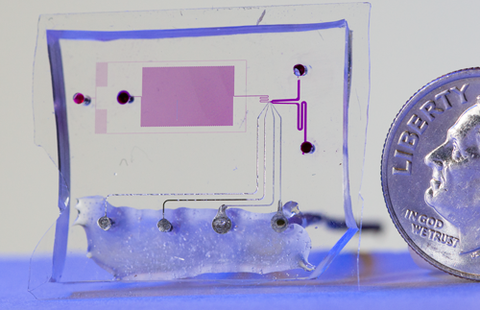
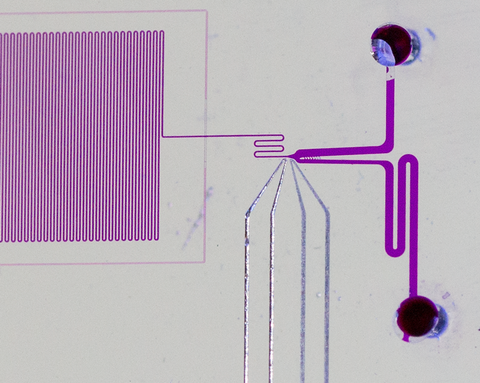
To do so, Jimenez's research group had to devise a one-of-a-kind technology: a chip-based microfluidic channel system that examines a single-file stream of cells containing different RFPs. As each cell flows down a channel a few tens of micrometers wide, it is exposed to multiple pulses of laser light and the RFP's emission is measured.
By comparing the intensity of light emitted after the first and last pulses, the scientists can see which cells are least susceptible to bleaching. And by comparing the phase difference between the laser pulse and the emitted red light, they can estimate the cell's fluorescence lifetime.
The process takes about 10 milliseconds per cell. During that time, electronics process the signals and determine which cells have superior characteristics. That triggers a tiny infrared tractor beam that pushes the best performers into a different channel where they are saved. The rest proceed down the main channel and are discarded. (See animation above.)
The research necessarily involves multiple iterations, in which a subset of the best performers from one generation is selected and mutated for the next generation. The process is repeated over and over in search of optimal characteristics.
"We can analyze thousands in an afternoon," Jimenez says, "and because the selected cells are alive when they come out of our channel, we can grow them up, examine their genes, and compare the biophotonic characteristics of the encoded RFPs."

But there is much more to learn as the researchers search for the best variations on the structure of the protein.
A typical RFP is a barrel-shaped arrangement of some 270 amino acids folded in a sort of basket-weave pattern. In the center of the barrel is a chromophore – the substance that emits red light after being excited. One way that chromophores degrade is by contact with oxygen, a highly reactive element. But animals require oxygen to live, and it permeates cells.
"So we need to know how to modify the structure of these things so that they produce more emission even in the presence of oxygen," Jimenez says. "We know that the protein barrels have holes in them through which oxygen is transmitted. We're looking for mutations in which those holes are partly or even entirely closed, or for ways to modify the gene to achieve the same effect. If we could eliminate the ability of oxygen to get in, if the barrel were tighter, we could get much better performance. That's an active area of research in our group."
"We're not looking for a single needle in a haystack, the one RFP that everybody will want," Jimenez says. "There is no silver bullet. Different experiments call for different RFP characteristics in different environments. Our aim is to create whole libraries of variants on red fluorescent proteins that can become the world standard for use in cellular imaging."
*Fluorescence is the process in which objects spontaneously emit light after absorbing photons from an outside source, akin to the phenomenon that makes luminous dials glow on clocks and watches. Fluorescent proteins were first identified in the early 1960s as part of the system that produces bioluminescence in marine organisms. The genes for such proteins can be easily transferred into other kinds of cells, and manipulated such that they are genetically fused to other proteins. Fluorescence doesn't affect the functions of those proteins.

