Taking Measure
Just a Standard Blog

Bad Blood
Four years ago, I found myself in a doctor’s office for an aging-related problem I started experiencing. To ease my anxiety, my doctor initiated some small talk by asking what I do for a living.
“I am a scientist studying a molecule called a carbon nanotube. It is made of a layer of carbon atoms that are arranged like a tube,” I replied. “It is geometrically beautiful, and it can potentially be used for many things.” I became a little bit excited and told him an idea my collaborators and I had just come up with recently: “One of them is to analyze blood for diagnosing any kind of disease!” My doctor did not seem impressed. He smiled politely, told me that he had just read the nonfiction book Bad Blood, and recommended that I read it too.
I took my doctor’s advice and read the book. It told the story of a failed Silicon Valley startup that claimed to be able to diagnose multiple diseases using the blood from just a pinprick. The founder, Elizabeth Holmes, certainly got it right that blood-based diagnosis has huge potential. But it was obvious to me that she wouldn’t be able to make it by just repackaging existing technologies. I lamented the fact that Holmes was so successful in attracting investment while my collaborators and I had such a hard time getting funding for our idea.
A Tale of Two Molecules
Unlike Holmes, we had been working for decades on new biosensing materials and methods.
In 2002, while working at DuPont Central Research and Development, my colleagues and I found that a chain of DNA — the molecule of life — can wrap around a carbon nanotube to form a new hybrid molecule. Since then, we and others have found that this hybrid molecule has many interesting properties.
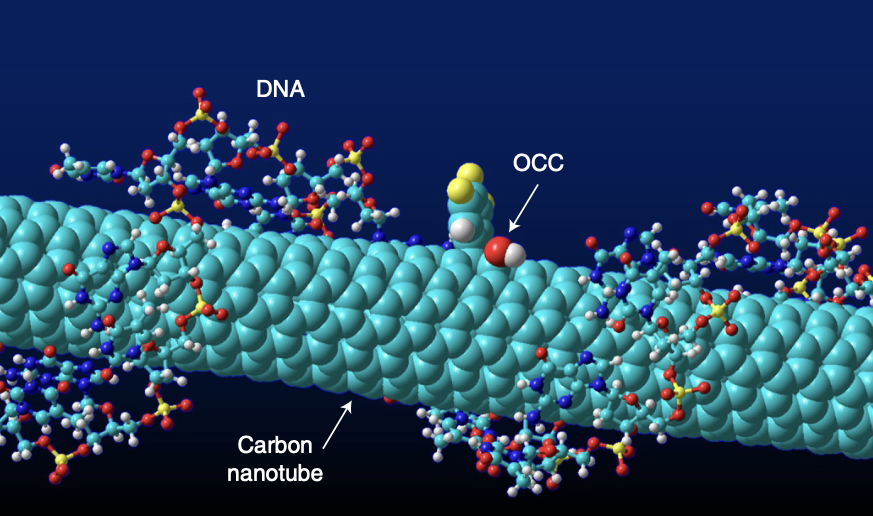
One of them is that the hybrid molecule is exquisitely sensitive to the biomolecules around it. This is because all the atoms of a carbon nanotube are on its surface, making it ideally suited for sensing its chemical environment. A layer of DNA coating on the surface provides a friendly and tunable interface between the nanotube and the biomolecules. By changing the DNA sequence and nanotube structure, we can build many sensors to detect different molecules. We could also cause the nanotube to emit light, or fluoresce, based on what it was sensing. How could we make use of this unlimited, easily accessible molecular resource?
Mimicking Perception
We thought that a good way to use this resource would be to analyze something as complex as blood. Blood contains thousands of molecules, with concentrations spanning several orders of magnitude. A blood test provides lots of information to your doctor regarding your physiological state, but a definitive diagnosis of certain disease is often difficult because it may involve subtle changes in the concentrations of many different molecules.
Differentiating disease from healthy conditions by testing blood can be compared to differentiating two wines that have subtle differences in their chemical composition. Experienced sommeliers do this by relying on their sense of perception, using a plethora of taste buds and years of experience stored in their brain. Why not mimic this process to diagnose disease by “tasting” (or sensing) blood with DNA-carbon nanotube sensors and interpreting the data using artificial intelligence? I call this scheme of disease detection “molecular perceptron” for its perception-mimicking principle and for the cool suffix “-tron” we find in terms like “synchrotron,” “cyclotron,” “neutron,” etc.
Detecting Ovarian Cancer
In 2019, my collaborators (Dan Heller of Memorial Sloan Kettering Cancer Center, Anand Jagota of Lehigh University and YuHuang Wang of University of Maryland) and I decided to give the idea a try despite the lack of any direct funding. Dan is a leading expert on biosensing, Anand and I have worked on DNA-carbon nanotube hybrid molecules for nearly two decades, and YuHuang is an expert on carbon nanotube chemistry. We decided to test blood samples for ovarian cancer. This is a deadly disease, and earlier detection and timely measurement of disease progression and recurrence would markedly improve outcomes for patients.
Mijin Kim, a talented postdoctoral fellow in Dan's lab, took the lead and pursued the project with great enthusiasm. The current procedure for diagnosing ovarian cancer involves measuring the concentration of a single blood protein and conducting a physical examination using transvaginal ultrasonography.
After two years of hard work, we showed that our method can diagnose ovarian cancer more accurately than the current procedure. For this and other accomplishments, Mijin was recently recognized as one of MIT Technology Review’s 2022 “Innovators Under 35.”
Turning Science Fiction Into Reality
We are all excited by this initial success. What is more exciting is the future prospect of our approach.
“To see a world in a grain of sand” is one of my favorite lines by William Blake. What we are trying to accomplish is to see life in a drop of blood!
Our molecular perceptron design is not specific to any particular medical condition. It should be able to detect a vast number of diseases and provide a comprehensive picture of physiological condition in general. And like human learning, it will get better and better with more data.
We envision that in the future, every human being on Earth will have their molecular perceptron data recorded periodically to track physiological changes and to guide preventive measures. I want to do as much as I can to turn our vision into reality: design sensor elements, calibrate the sensors, establish standards and more. I view this as doing my job “in ways that … improve our quality of life,” matching exactly the final words of NIST’s mission statement.






Bravo!!