Summary
In this project, we investigate appropriate uncertainty analysis and splicing algorithms for spectral data, especially fluorescence spectroscopy calibration, to support bioassay standard development efforts.
Description
Biological assays are procedures that can determine concentration, purity, or biological activity of a substance through scientific experimentation. Fluorescence-based quantification one of the most widely used technique in quantitative clinical and biochemical assays, and it depends on the measurement of relative intensities of fluorescence with wavelength. Fluorescence spectroscopy yield absolute signals, i.e., they are not ratios like absorbance measurements. Fluorescence Spectroscopy instrument has a unique spectral responsivity making both the spectral shape and absolute intensity of a single sample different on every instrument and even on a single instrument at different times. Thus, developing relative intensity standards for fluorescence instrument calibration serves a critical need of the biochemical community to achieve repeatable and reproducible measurement results. For the past several years we have developed uncertainty analysis in support of developing relative intensity standards for calibration of fluorescence spectroscopy. The goal of this project is to document the novel statistical methods and to disseminate related algorithms /software to the wide user community.
Major Accomplishments
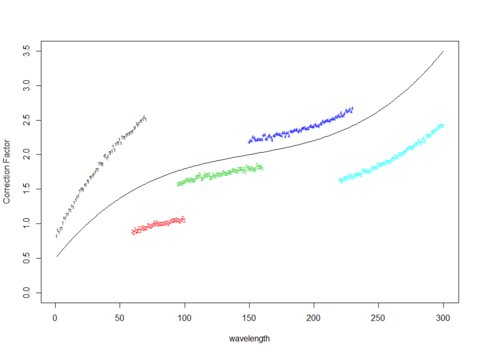
We have developed statistical methods for certifying spectrum-like quantities using analysis of variance for functional type data, and have developed a splicing algorithm for combining curves, or connecting dots, from different SRM calibration curves which can differ in scales but having very little overlap in the wavelengths being measured.

