Summary
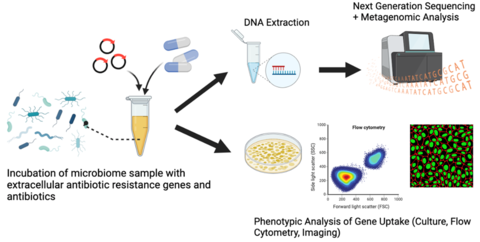
NIST is developing microbial measurements to better understand the propagation of antimicrobial resistance (AMR) through a variety of mechanisms. This work includes an exploration of horizontal gene transfer (from extracellular genes to cells) and vertical gene transfer (microbial evolution). NIST scientists have also explored the surveillance of antimicrobial systems through wastewater systems across the globe.
Description
NIST scientists are leveraging a wide range of measurement capabilities to study microbial community dynamics with regard to antimicrobial resistance. Using a range of culture, molecular and sequencing methods, we hope to develop methods to better understand antibiotic resistance evolution, transfer, and overall prevalence. In doing so, NIST scientists aim to provide a roadmap of microbial measurements used to track changes within complex and evolving microbiomes and gather valuable research findings on a pressing environmental and clinical issue.
Genetic evolution of the resistance profile within microbial communities via mutation (vertical gene transfer) and mobile genetic elements (horizontal gene transfer) is mapped using multiple genotypic and phenotypic measurements, including: CFU, genome copy number via PCR, relative and normalized taxonomic abundance via whole-genome sequencing, re-sequencing and variant calling, microscopic imaging, flow cytometry and gene expression via transcriptomic measurements.
NIST is focusing on three primary focus areas:
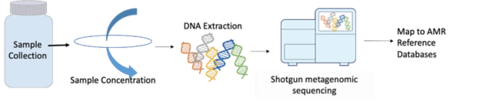
Sequencing strategies for AMR surveillance
Vertical Gene Transfer

Horizontal Gene Transfer