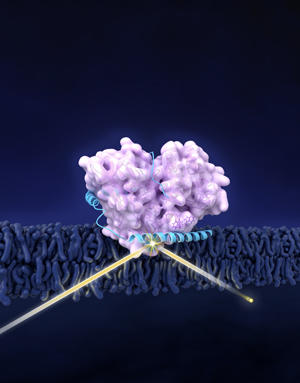
The proteins GCase (in pink) and α-syn (blue) forms a complex in cellular membrane. Neutron reflectometry (suggested by the yellow beam) revealed the structure of the complex. α-Syn shifts GCase slightly away from membrane, possibly contributing to effects related to Parkinson’s disease.
To understand diseases like Parkinson's, the tiniest of puzzles may hold big answers. That's why a team including scientists from the National Institute of Standards and Technology (NIST) have determined how two potentially key pieces of the Parkinson's puzzle fit together, in an effort to reveal how the still poorly understood illness develops and affects its victims.
This puzzle is a tough one because its pieces are not only microscopic but three-dimensional, and can even change shape. The pieces are protein molecules whose lengthy names are abbreviated as GCase and α-syn. The two proteins wrap around each other and take on a complicated shape before attaching themselves to the membrane surface inside a neural cell in a victim's brain.
While much remains unknown about Parkinson's, clues abound that the proteins' behavior is somehow important. Parkinson's victims have a buildup of α-syn in their cells, a possible factor in the dementia that the disease often brings. They also are far more likely to have a mutation in the gene that instructs cells to create GCase. Low levels of GCase cause another disease, Gaucher, and in some individuals suffering from both Parkinson's and Gaucher simultaneously, Parkinson's may appear at a younger age.
To get a better handle on how these proteins operate in the body, the team—which also included scientists from the National Institutes of Health (NIH) and Carnegie Mellon University—came to the NIST Center for Neutron Research (NCNR) to get a picture of how the two proteins combine into a single unit called a complex that interacts with cell membranes. Using techniques including neutron reflectometry, the team teased out the first-ever structural picture of the GCase/α-syn complex, including their shape change, which NIH's Jennifer Lee says would not have been detectable by any other methods.
"It gives us a potential interaction model of the two and structural insights in how α-syn may interfere with activity on a cell membrane," Lee says. "An equally important contribution here is that this is the first, I believe, to look at complex formation at the membrane interface using neutron science."
The study still leaves many mysteries about the complex—notably how it attaches to and interacts with the membrane inside a part of the cell called the lysosome where α-syn is broken-down. The team plans to follow up with additional investigations, especially once an improved reflectometer that could offer greater resolution arrives at the NCNR in about 2017. Meanwhile, the results of the current study—and the tools that provided them—will give the team other options to explore.
"It lays the groundwork for exploring the complex relationship between proteins that are involved in the causes of disease," Lee says. "This will also offer a way for us to investigate other substances that would affect the interaction."
*T.L. Yap, Z. Jiang, F. Heinrich, J.M. Gruschus, C.M. Pfefferkorn, M. Barros, J.E. Curtis, E. Sidransky and J.C. Lee. Structural features of membrane-bound glucocerebrosidase and α-synuclein probed by neutron reflectometry and fluorescence spectroscopy. Journal of Biological Chemistry, DOI: 10.1074/jbc.M114.610584, Jan. 9, 2015.

