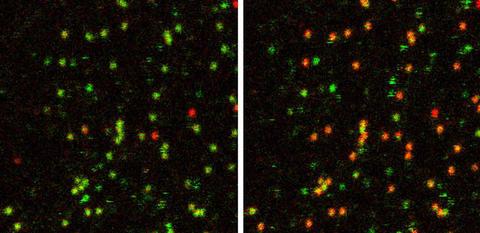
These false-color images show the behavior of about 30 to 40 single RNA molecules tagged with fluorescent dyes in the absence of magnesium (left) and with high magnesium concentrations (right). Green indicates that the tagged molecules are farther apart (undocked) whereas red indicates they are closer together (docked), showing that magnesium promotes docking.
Biophysicists have developed a method for studying, in real time, a nanoscale “docking and undocking” interaction between small pieces of ribonucleic acid (RNA), a technique that may be broadly useful in studying structural changes in RNA that affect its function. The research at JILA, a joint institute of the National Institute of Standards and Technology (NIST) and University of Colorado at Boulder, may have applications in the design of effective new drugs based on small RNA strands.
RNA is a chain-like molecule that contains genetic information, makes proteins and catalyzes biological reactions. Scientists at JILA are studying RNA using methods that reveal how individual chemical units of RNA dock, or lightly and temporarily bond, to form special three-dimensional shapes that exhibit biochemical activity. The latest work, to be published the week of July 11 in the Proceedings of the National Academy of Sciences,* adds to understanding of the intramolecular “stickiness” between specific loops and sequences in the RNA that help stabilize this folding. This type of information is crucial to understanding RNA structure and, ultimately, how it affects function.
The JILA group developed a simple model system for studying the reversible docking of a small piece of RNA at a receptor site in the same molecule. They used a technique called fluorescence resonance energy transfer, in which the two pieces of RNA are labeled with different dyes that have overlapping emission bands. One dye emits light of the same color that the other dye absorbs; the second dye then emits light of a different color. One piece of RNA is excited by a laser and, when the two pieces are close enough together to dock, passes energy to the other one, which then fluoresces. This method was used to measure the distance between the two pieces of RNA as it varied from less than 4 nanometers in the docked state to about 7 nm in the undocked state.
The scientists used ultrasensitive laser-based microscopy methods to image many isolated RNA molecules simultaneously, in effect generating a “movie” of single molecule docking kinetics in real time. They used this method to study thousands of pieces of RNA over time scales of 10 to 30 seconds, and observed about two-thirds of them rapidly docking and undocking. The rates of docking and undocking were measured as a function of the concentration of magnesium ions in the surrounding fluid, revealing a complex dependence on metal ions, as is typical for RNA. The docking rate rose 12-fold as magnesium concentrations increased. A significant number of molecules still docked in the absence of magnesium—the first time this phenomenon has been observed, according to the paper.
The research is supported by NIST, the National Science Foundation, National Institutes of Health, and the W.M. Keck Foundation initiative in RNA science at CU-Boulder.
*J.H. Hodak, C.D. Downey, J.L. Fiore, A. Pardi and D.J. Nesbitt 2005. Docking kinetics and equilibrium of a GAAA tetraloop-receptor motif probed by single molecule fluorescence resonance energy transfer. Proceedings of the National Academy of Sciences. Week of July 11.

