Summary
This project applies hydrogen/deuterium exchange mass spectrometry (HDX-MS) for determining the dynamical structure of therapeutic antibodies, glycoproteins, and proteins; transmembrane protein receptors; and ligand-receptor interactions. Topics of particular interest include protein-ligand interactions, structure-function relationships between glycan structure and glycoprotein folding, and comparability measurements among innovator drugs and biosimilar drug candidates. This project aims to improve measurements through method and instrumentation development, to validate HDX-MS measurement technology by conducting interlaboratory comparisons, and to improve measurement accuracy through parallel HDX-MS and HDX-NMR studies of selected proteins and glycoproteins.
Description
Intended Impact
This project is designed to improve, test, and validate hydrogen/deuterium exchange mass spectrometry (HDX-MS) metrology for the determination of dynamical properties of therapeutic proteins and glycoproteins.
Objectives
- Explore structure-function relationships between glycan structure and protein folding energies
- Develop HDX-MS for assessments of dynamical comparability among innovator and candidate biosimilar drugs
- Measure the reproducibility of proteolytic fragmentation HDX-MS by conducting interlaboratory comparisons on a Fab protein
- Improve HDX-MS technology for measurements of transmembrane protein drug targets
- Measure interactions of biological drugs with surfaces and aggregates
- Enhance resolution of the HDX-MS method to the single amide level
- Adopt internal H/D exchange reporters and materials to become NIST standard reference materials for improvement of reproducibility and harmonization of HDX-MS
Technical Approach
The research project uses HDX-MS to measure the D-for-H exchange rates of the amide groups along the backbone of a protein in D2O solution. These rates indicate the protection factors of the amide groups. The protection factors are characteristic of higher order structural features of proteins (e.g., α-helixes, β-sheets ...) that are stabilized through hydrogen bonding, disulfide bonds, electrostatic interactions, and hydrophobic forces. Since protection factors can change as the protein binds with ligands or undergoes folding, HDX-MS can provide sensitive diagnostic data evidencing structural differences. HDX-MS is a rapidly evolving metrology. Numerous improvements in automation, instrument resolution and accuracy, and new, more powerful software are being introduced.
Major Accomplishments
HDX-MS Interlaboratory Comparison Project
An interlaboratory study involving 37 co-authors in 15 unharmonized laboratories determined the reproducibility of continuous-labeling, bottom-up HDX-MS measurements by measuring the deuterium uptake of Fab of NISTmAb (PDB: 5K8A) for immersions in D2O for tHDX = (30 to 14400) s and THDX = (25, 21, 3.6) oC. Each laboratory was sent a standardized kit containing Fab protein the reagents necessary to conduct experiments.
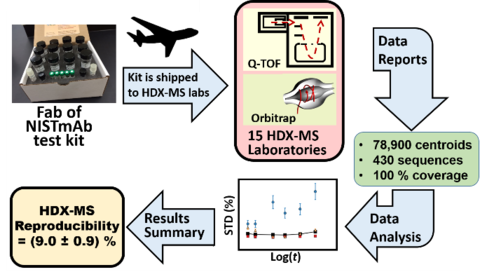
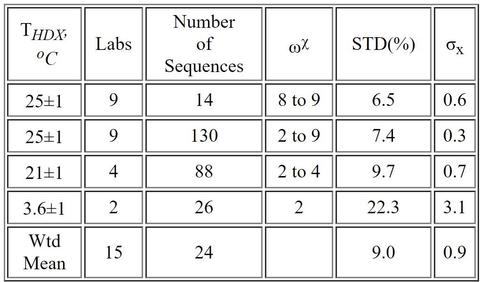
Laboratories reported ~ 89,800 centroid measurements for 430 proteolytic peptide sequences of the Fab fragment (~78,900 centroids), giving nearly 100 % coverage, and approximately 10,900 centroid measurements for 77 peptide sequences of the Fc fragment. Nearly half of peptide sequences are unique to the reporting laboratory, and only two sequences are reported by all laboratories. The majority of the laboratories (87 %) exhibited centroid mass laboratory repeatability precisions of ≤ (0.15 ±0.01) Da (1σx). All laboratories achieved ≤ 0.4 Da. For immersions of protein at THDX = (3.6 to 25) oC and for D2O exchange times of tHDX = (30 s to 4 h) the reproducibility of back-exchange corrected, deuterium uptake measurements for the 15 laboratories is (9.0 ±0.9) % (1s). A 9 laboratory cohort that immersed samples at THDX = 25 oC exhibited reproducibility of (6.5 ±0.6) % for back-exchange corrected, deuterium uptake measurements. (See refs 2, 3 below.)
Automated Removal of Phospholipids from Membrane Proteins
NIST researchers have developed a new, automated approach to study the structural dynamics of membrane proteins, the most common targets for pharmaceuticals. The new method involves isolation of proteins from their phospholipid bilayer using a scheme that is compatible with robotic handling. The key development was the introduction of ZrO2 microbeads, which strongly coordinate with the membrane phospholipids, and then removal of microbead-coordinated phospholipids by inline nanofilters. The protein is thus freed from phospholipids, which interfere with analysis by LC-MS.
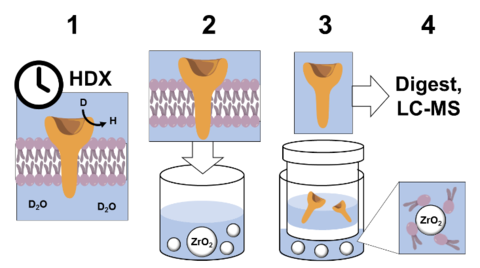
Previous HDX-MS publications on membrane protein dynamics removed phospholipids manually, requiring arduous, hours-long laboratory work. The new approach was demonstrated on the transmembrane protein FcγRIIa, and showed information about the regions outside of, across, and inside of its liposomal membrane, enabling measurement of structural dynamics of the whole protein. This approach is the first to enable fully automated HDX-MS on full-length transmembrane proteins in lipid bilayers. (See ref. 4 below.)
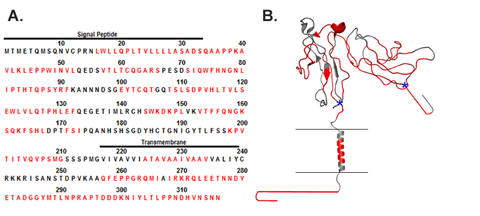
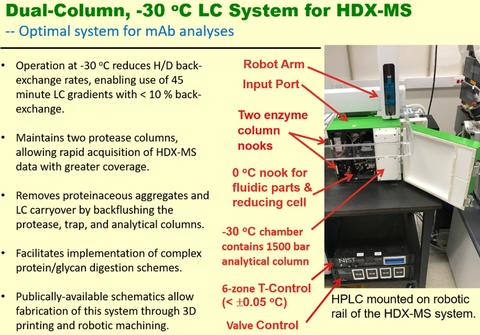
HDX-MS Subzero Analysis System
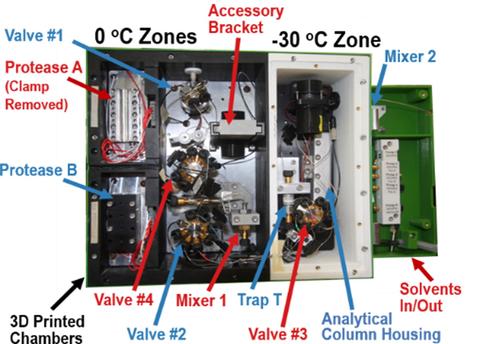
New applications for HDX-MS, particularly those involving quality control and biosimilarity evaluations, will demand deuterium uptake measurements of improved precision. NIST researchers have developed an analysis system for HDX-MS that conducts chromatography at ‑30 °C. Extending gradients to 40 minutes at -30 °C exhibited ≈16 % more deuterium than for a conventional 8 minute gradient at 0 °C, and a subset of peptides showed ≈26 % more deuterium. Chromatographic resolution is improved, allowing HDX-MS measurements on a greater number of peptides. Residual back-exchange of ≤ 10 % still occurs, as this is governed by the sample preparation process at 0 °C. Total measurement uncertainty is greatly reduced. Reproducibility is enhanced by regulating chamber temperatures to ±0.05 °C. Isotopic bias is effectively eliminated by executing complex cleaning cycles that include back-flushing of the proteolytic and chromatographic columns. To increase the diversity of proteolytic peptides, the analysis system stores two protease columns that are perpetually cleaned and always ready for use.
Features of the subzero (-30 oC) protease and chromatography system.


Other Accomplishments
-
Demonstrated that HDX-MS can detect dynamical differences among glycoforms of the same glycoprotein. (ref. 9)
-
Developed a method that measures D-uptake by glycans within glycoproteins (ref. 9).
-
Used HDX-MS and mutagenesis to find the epitope of Proliferating Cell Nuclear Antigen (PCNA) and a previously unknown ligand, TIP. This ligand offers a new vector for attacking solid tumor cancers (ref. 10).
Publications
- "Chromatography at -30 °C for Reduced Back-Exchange, Reduced Carryover, and Improved Dynamic Range for Hydrogen-Deuterium Exchange Mass Spectrometry" K. W. Anderson, J. W. Hudgens, Journal of the American Society for Mass Spectrometry 33(7), 1282-1292 (2022). DOI: 10.1021/jasms.2c00096.
- "Interlaboratory Studies Using the NISTmAb to Advance Biopharmaceutical Structural Analytics" K. Yandrofski, T. Mouchahoir, M. De Leoz, D. Duewer, J. W. Hudgens , K. W. Anderson, L. Arbogast, F. Delaglio , R. G. Brinson, J. P. Marino , K. Phinney, M. Tarlov , J. E. Schiel, Frontiers in Molecular Biosciences 9, 876780 (2022). DOI: 10.3389/fmolb.2022.876780.
- "HDX-MS and MD Simulations Provide Evidence for Stabilization of the IgG1-FcγRIa (CD64a) Immune Complex Through Intermolecular Glycoprotein Bonds" K. W. Anderson, C. Bergonzo, K. Scott, I. L. Karageorgos, E. S. Gallagher, V. S. Tayi, M. Butler, J. W. Hudgens, Journal of Molecular Biosciences 434(2), 167391 (2022). DOI: 10.1016/j.jmb.2021.167391.
- "Dataset from HDX-MS Studies of IgG1 Glycoforms and Their Interactions with the FcγRIa (CD64) Receptor" K. W. Anderson, K. Scott, I. L. Karageorgos, E. S. Gallagher, V. S. Tayi, M. Butler, J. W. Hudgens, Journal of Research of the National Institute of Standards and Technology;126010 (2021). DOI: 10.6028/jres.126.010.
- "Conformational gating, dynamics and allostery in human monoacylglycerol lipase" S. Tyukhtenko, X. Ma , G. Rajarshi , I. Karageorgos, K. W. Anderson, J. W Hudgens, J. J. Guo, M. L. Nasr , N. Zvonok , K. Vemuri, G. Wagner, A. Makriyannis, Scientific Reports 10(1), 18531 (2020). DOI: 10.1038/s41598-020-75497-5.
- "Construction of a Dual Protease Column, Subzero (-30 oC) Chromatography System and Multi-channel Precision Temperature Controller for Hydrogen-Deuterium Exchange Mass Spectrometry" J. W. Hudgens, Journal of Research of the National Institute of Standards and Technology 125, 125025 (2020).
DOI: 10.6028/jres.125.025, Database Archive: https://doi.org/10.18434/M32151 - "Interlaboratory Comparison of Hydrogen–Deuterium Exchange Mass Spectrometry Measurements of the Fab Fragment of NISTmAb"
J. W. Hudgens, I. Karageorgos, E. S. Gallagher, K. W. Anderson, A. Espada, M. J. Chalmers, E. Harguindey, H-M Zhang, B. Walters, J. Zhang, J. Venable, C. Steckler, I. H. Park, A. Brock, X. Lu, R .H. Pandey, A. Chandramohan, G. S. Anand, S. N. Nirudodhi, J. Sperry, J. Rouse, J. A. Carroll, K. D. Rand, U. Leurs, D. D. Weis, M. A. Al-Naqshabandi, T. S. Hageman, D. Deredge, P. Wintrode, M. Papanastasiou, J. D. Lambris, S. Li, and S. Urata, Analytical Chemistry 91, 7336-7345 (2019). DOI: 10.1021/acs.analchem.9b01100 - "Hydrogen-Deuterium Exchange Mass Spectrometry (HDX-MS) Centroid Data Measured between 3.6 oC and 25.4 oC for the Fab Fragment of NISTmAb" J. W. Hudgens, I. Karageorgos, E. S. Gallagher, K. W. Anderson, A. Espada, M. J. Chalmers, E. Harguindey, H-M Zhang, B. Walters, J. Zhang, J. Venable, C. Steckler, I. H. Park, A. Brock, X. Lu, R .H. Pandey, A. Chandramohan, G. S. Anand, S. N. Nirudodhi, J. Sperry, J. Rouse, J. A. Carroll, K. D. Rand, U. Leurs, D. D. Weis, M. A. Al-Naqshabandi, T. S. Hageman, D. Deredge, P. Wintrode, M. Papanastasiou, J. D. Lambris, S. Li, and S. Urata, Journal of Research of the National Institute of Standards and Technology 124, 124009 (2019) DOI: 0.6028/jres.124.009, Database Archive: DOI: 10.18434/8SX3-NQ49
- "Automated removal of phospholipids from membrane proteins for H/D exchange mass spectrometry workflows" K. W. Anderson, E. S. Gallagher, and J. W. Hudgens, Analytical Chemistry 90, 6409−6412 (2018). DOI: 10.1021/acs.analchem.8b00429
- “Effects of Distal Mutations on the Structure, Dynamics and Catalysis of Human Monoacylglycerol Lipase” S. Tyukhtenko, G. Rajarshi, I. Karageorgos, N. Zvonok, E. S. Gallagher, H. Huang, K. Vemuri, J. W. Hudgens, X. Ma, M. L. Nasr, S. Pavlopoulos, and A. Makriyannis, Scientific Reports 8, 1719 (2018). DOI: 10.1038/s41598-017-19135-7
- "In vitro cytochrome P450 46A1 (CYP46A1) activation by neuroactive compounds" N. Mast, K. W. Anderson, K. M. Johnson, T. Phan, F. P. Guengerich, I. A. Pikuleva, Journal of Biological Chemistry 292(31), 12934-12946. DOI: 10.1074/jbc.M117.794909.
- "Conformational Changes in Active and Inactive States of Human PP2Cα Characterized by Hydrogen/Deuterium Exchange-Mass Spectrometry" S. J. Mazur, E. S. Gallagher, S. Debnath, S. R. Durell, K. W. Anderson, L. M. M. Jenkins, E. Appella, and J. W. Hudgens,
Biochemistry, 56, 2676−2689 (2017). DOI: 10.1021/acs.biochem.6b01220. - "Mapping of the Allosteric Site in Cholesterol Hydroxylase CYP46A1 for Efavirenz, a Drug That Stimulates Enzyme Activity." K. W. Anderson, N. Mast, J. W. Hudgens, J. B. Lin, I. V. Turko, and I. A. Pikuleva, Journal of Biological Chemistry, 291, 11876-11886 (2016). DOI:10.1074/jbc.M116.723577
- "Mapping Protein–Ligand Interactions with Proteolytic Fragmentation, Hydrogen/Deuterium Exchange-Mass Spectrometry" E. S. Gallagher and Jeffrey W. Hudgens, Methods in Enzymology,566, 357-404. (2016). DOI: 10.1016/bs.mie.2015.08.010
- "The Effects of Desialylation on Human α1-Acid Glycoprotein-Ligand Interactions" R. Y.-C. Huang and J. W. Hudgens, Biochemistry, 52, 7127-7136 (2013). DOI: 10.1021/bi4011094
- "A Novel mechanism for regulating the activity of proliferating cell nuclear antigen by a small protein" Z. Li, R. Y.-C. Huang, D. C. Yopp, T. H. Hileman, T. J. Santangelo, J. Hurwitz, J. W. Hudgens, and Z. Kelman, Nucleac Acids Research, 42(9), 5776-6789 (2014). DOI: 10.1093/nar/gku239
- Book Chapter: "Method Validation and Standards in Hydrogen Exchange Mass Spectrometry" J. W. Hudgens, R. Y.-C. Huang and E. D'Ambro, in Hydrogen Exchange Mass Spectrometry of Proteins: Fundamentals, Techniques and Applications; David Weis, ed. (Wiley, 2016).
- Book Chapter: "Emerging Technologies to Assess the Higher-Order Structure of Monoclonal Antibodies" J.P. Marino, R.G. Brinson, J.W. Hudgens, J.E. Ladner, D.T. Gallagher, E. S. Gallagher, L.W. Arbogast, and R.Y.-C. Huang, In Emerging Technologies To Assess the Higher Order Structure of Monoclonal Antibodies. In State-of-the-Art and Emerging Technologies for Therapeutic Monoclonal Antibody Characterization Volume 3. Defining the Next Generation of Analytical and Biophysical Techniques, Schiel, J.; Borisov, O.; Davis, D., eds. American Chemical Society: 2015; Vol. 1202, pp 17-43. DOI:10.1021/bk-2015-1202.ch002

