Summary
A collaboration between the Medical University of South Carolina (MUSC), Hollings Marine Laboratory, and NIST is examining the relationship between cell phenotype and gene expression in lung endothelial cells from dolphin (Tursiops truncatus). The overall goal of this research collaboration is to lay the groundwork for the metrology infrastructure required for cell and tissue-based screening of marine mammal health. The initial aim is to compare morphological and other image-derived data to gene expression profiles of bottlenose dolphin lung cells.
Description

Due to their characteristics (long life span, feed at a high trophic level, extensive fat stores that can act as a deposit for anthropogenic toxin) dolphins have been proposed as a good sentinel organism in aquatic and coastal environments to monitor and understand the impact of infectious diseases (toxoplasmosis, urogenital cancer), anthropogenic pollutants (organohalogen and heavy metal contaminants), and harmful algal blooms and their toxins (brevetoxins, saxitoxins, okadaic acid). In this collaborative work, the physiology of a wild animal, the dolphin, was investigated. Dolphin lung endothelial cells were used to investigate the unique physiological adaptation of the dolphin lungs that could have therapeutic application to human related respiratory disease (fourth leading cause of death in the United States). The dolphin lung cells were challenged in this study with apoxic (low oxygen) conditions and with various toxins.
This project is intended to lay the groundwork for the metrology infrastructure required for cell and tissue-based screening of marine mammal health. The initial portion of this project aims to comprehensively characterize the growth characteristics of the dolphin lung cells. Future efforts will focus on comparison of morphological and image-derived data with dolphin lung cell gene expression profiles, particularly in response to apoxic conditions and toxin treatments.
Primary dolphin lung endothelial cells from a fetal calf were provided by the University of Mississippi, and were single-cloned at passage 13 to select for the lung endothelial cells. Cells in culture were quantitatively compared to bovine lung enodothelial cells, human umbilical vein endothelial cells, and rat fibroblasts, with respect to morphological characteristics, such as spread area, roundness, aspect ratio, and tendency to maintain cell-cell contacts. These measurements were made by fixing and staining cells and collecting digitized images using fluorescence microscopy. Image analysis was performed with open source software, ImageJ.
The characterization of the dolphin cells was based on the analysis of four cultures of the cells from different frozen stocks cultured over ~ 50 passages. To further describe the cell line characteristics specific markers, including PECAM-1, VE-Cadherin, and Ac-LDL uptake were used to investigate the endothelial nature of the cells through immunostaining and Western blotting.
The growth rates and division times for these cell cultures were determined from their characteristic distributions of cell volumes, measured by impedance and the application of a mathematical model of growth and division developed by Michael Halter and John Elliott.
RNA was also collected every other passage fur further genomic analysis.
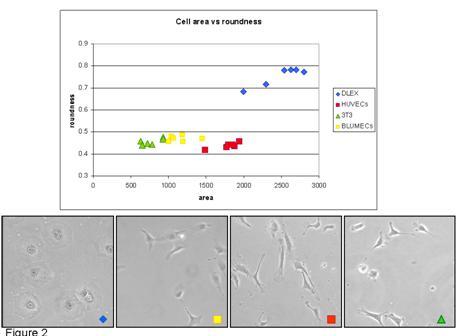
Phase microscopy of (blue) dolphin cells, (yellow) human umbilical vein endothelial cells, (red) rat fibroblasts, and (green) bovin lung endothelial cells. Quantitation of cell roundness for the four cell types is shown in the graph.
Major Accomplishments
- Dolphin cell volumes in culture were observed to decrease over time, suggesting more rapid division rates. This change was accompanied by corresponding changes in cell volume and, often, changes in morphology.
- Although the dolphin cells do not show positive responses to the typical endothelial markers (which could also be due to a limitation of species-specific antibodies), the morphological changes observed are consistent with previously observed trans-differentiation events (e.g. endothelial-to-mesenchimal transition). This observation will likely be of great significance to endothelial cell biology of many species in addition to dolphin.
- Publication: Quantitative methods to characterize morphological properties of cell lines

