Image segmentation based measurements
Summary:
The problem of segmentation, when cells are in contact, poses a challenge to the accurate automation of cell counting, tracking and lineage modeling in cell biology. The segmentation methods used consist of:
- background reconstruction to obtain noise-free foreground pixels, and
- incorporation of biological insight about dividing and non-dividing cells into the segmentation process to achieve reliable separation of foreground pixels into pixels associated with individual cells.
The segmentation results for a time lapse image stack were compared against manual segmentation provided by experts, which we consider as reference data. We chose two metrics to measure the accuracy of segmentation:
- the Adjusted Rand Index (ARI) which compares similarities at a pixel level between masks resulting from manual and automated segmentation, and
- the Number of Cells per Field (NCF) which compares the number of cells identified in the field by manual vs. automated analysis.
Our results show that the automated segmentation compared to manual segmentation has an average ARI of 0.96 (1 being a perfect match), with a standard deviation of 0.03, and an average difference of the two NCF quantities equal to 5.39% with a standard deviation of 4.6% (n=238 images from a time lapse image sequence for both the ARI and NCF calculations).
Live cell tracker demo
Challenge Problem(s):
Automated segmentation and tracking pose significant challenges due to the variability of image and object qualities. The variability arises from differences in imaging devices and modalities. Additionally, different cell lines can have different physical characteristics and these characteristics can change in response to external conditions during a cell cycle. A particular challenge is accurately segmenting individual cells when cells are in contact with each other, particularly as the culture grows and cell density increases. The main motivation of this work is to address this problem and enable the collection of single cell data from cell monolayers as they approach confluence.
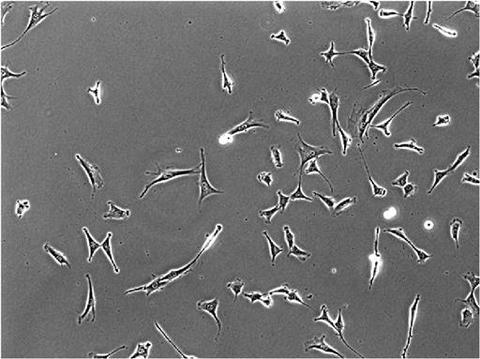
An example of a live cell phase contrast image of NIH 3T3 cells.
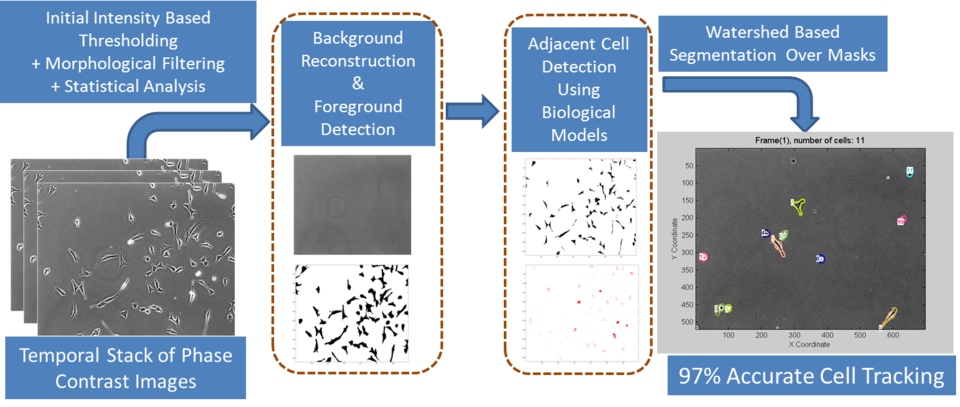
Overview of dynamic measurements in live cell workflow

