Image Quality and Classification Accuracy
Project research questions
- What microscopy conditions yield the best segmentation results?
- Which image quality descriptors relate best to segmentation accuracy?
- Which of the relevant descriptors require the least computational resources?
- How does consistency of reference data impact the performance of machine learning algorithms?
Project research challenges
- Wide range of cell lines, cell substrates, microscopes, and their settings makes it difficult to develop a generic solution.
- Inconsistent reference data degrades the accuracy of the machine learning methods.
Data Quality
To have confidence in the knowledge gleaned from data, it is essential to verify the quality of the data sources and to have a means of quantifying the potential uncertainties due to the quality of data for making critical, intelligent decisions. In computational biology, two areas where data quality is significant are quality of measured images and quality of the corresponding reference data (e.g., manual segmentations or cell colony labels).
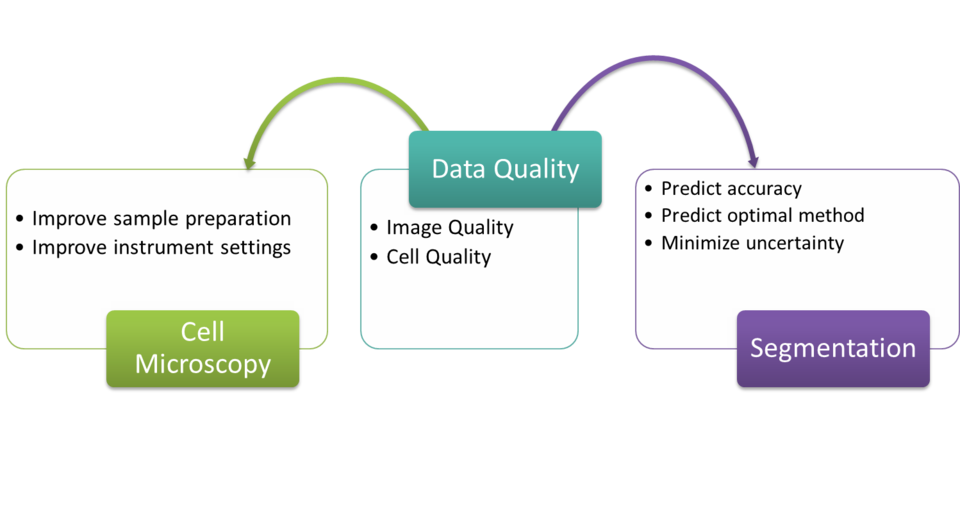
Research Focus
The objective of the data quality component in the CS-Bio-Met project is to collect and develop a repository of image quality descriptors and analyze their sensitivity in improving data quality upstream (microscope conditions, sample preparation) and downstream (recommending segmentation methods, predicting segmentation accuracy). Developing automated methods to assess image quality improves quality of the biological analysis. Automated image analysis ensures objectivity of the results. However the quality of the image directly affects the accuracy of the image analysis (segmentation) and has been proven to impact the accuracy of the research findings such as drug effectiveness and optimal dosage. This is true especially in applications such as the High Content Screening (HCS). HCS is an automated microscopy technique enabling the evaluation of spatial and temporal effects on cells for drug discovery and other applications. In addition to the quality of measured images, inconsistent reference labels that correspond to the measured images can degrade the ability to derive biologically meaningful classes or clusters. Inconsistent labels imply that the experts did not agree when determining the reference label, and hence, it is difficult to create rules for classification or clustering. The project aims at understanding the impact of reference data quality on clustering and classification uncertainty.

