Summary
As part of NIST's response to the Materials Genome Initiative (MGI), this project addresses the need for automation for the assembly and distribution of parallel simulation tasks.
Description
In computational materials science, many problems require the execution of numerous parallel simulation tasks on High Performance Computing (HPC) resources. Often a single published data point is the result of several parallel tasks executed in a specific sequence. Despite the continual improvement of computational capability, parallel simulation tasks are generally prepared, executed, and analyzed in a non-automated way via the command line and a job scheduler. If the cost savings and time reduction goals of the MGI are to be realized, automation is critical.
In response to the MGI, researchers in the Thermodynamics and Kinetics Group are investigating methods to automate the assembly and distribution of parallel simulation tasks, which we refer to as a scientific workflow. One obvious solution is the use of a traditional workflow management system. However, many traditional workflow management systems require wholesale changes in administrative and user activities. Therefore, we extend our investigation into non-traditional tools that can allow for incremental adoption and integration with existing HPC infrastructure.
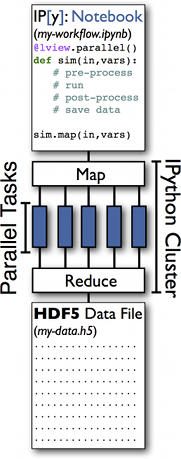
We are currently focused on the capabilities and limitations of IPython. We create an example workflow in IPython Notebook and distribute parallel simulation tasks using the IPython Cluster. The Notebook is an intuitive interface, which can be replaced by python code. The primary source of automation is the IPython Cluster, which is used as a dynamic gateway to HPC resources. IPython parallel allows simple and robust ways to distribute and execute tasks on the IPython cluster. Upon execution, the workflow distributes and executes all tasks to the appropriate IPython engines. In contrast, without automation, the user would manually manage the series of tasks, requiring significant time and with a much greater chance of error.
IPython has limitations and may not be a replacement for a traditional workflow management system in certain circumstances. Therefore, we continue to investigate scientific workflow tools that allow for the assembly and distribution of parallel simulation tasks.
Major Accomplishments
Using IPython, complete automation for the following simulations:
- Bulk mechanical properties
- Surface mechanical properties
- Generalized stacking fault energy

