Image conformance to algorithmic assumptions
Abstract:
Cell measurements are derived frequently from the results of pixel classification and contiguous region segmentation of microscopy images. Image segmentation is accomplished by classifying image pixels based on high dimensional image features computed from the pixel neighborhood. This poster addresses the conformance measurements of image features to Bayes classifier assumptions. Particularly, we focus on Gaussian probability distribution model of features and conditionally independent features. Our approach consists of three steps. First, we estimate multi-variate Gaussian model parameters from labeled data. Next, we generate synthetic data using modifications of the estimated parameters to obtain a fully conforming data set to each classifier assumptions. Finally, we compare the classification accuracies of Bayes classifiers with any combination of the two assumptions that are evaluated on labeled data and synthetic data (partially and fully conforming data to the assumptions). These conformance measurements are applied to 15 image features extracted from a set of sub-cellular fluorescent microscopy images of A10 fibroblast cells placed on soft or stiff extra cellular matrices (ECMs), and stained for actin, myosin, and focal adhesion. The classification accuracies are presented for segmentation of textured regions containing actin. Our conformance analyses of classifier assumptions yielded (a) uncertainty measurements, (b) a criterion for choosing the most suitable type of Bayes classifier, and (c) a cost function for selecting a reduced feature set satisfying classifier assumptions.
Description:
Problem Statement
Characterize classification error due to: Nonconformance to classifier assumptions Inability of the feature model and classifier to separate classes.
Objectives
Estimate uncertainty of the accuracy on the unlabeled dataset:
- Judge if training set is representative
- Estimate homogeneity/heterogeneity of dataset
- Select classifier & features for which data conforms to the assumptions.
Maximum Likelihood (ML) Gaussian model based Bayes Classifier:
- Features in the multi-dimensional space are normally distributed
- Features are also conditionally independent from each other's.
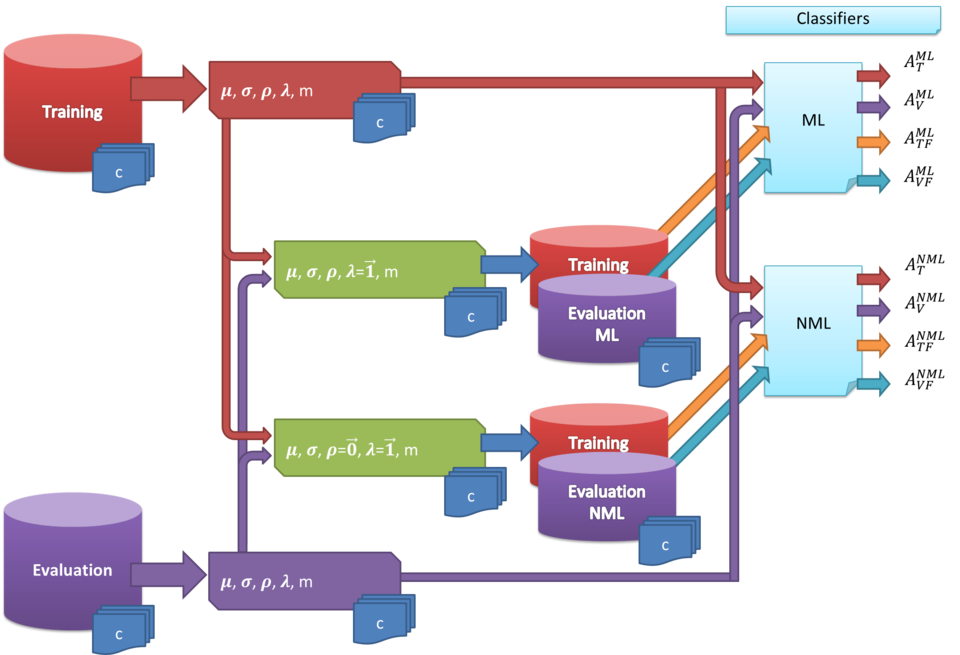
Approach Overview

Result Interpretation

