Summary
Mass spectrometry is an important technique in analytical chemistry, essential in areas including drug development, criminal and agricultural forensics, detection of explosives, and verification of chemical weapons treaty compliance. Yet there is no way to predict or model mass spectra from first principles. This means that a compound can only be identified with confidence if it is already known, with its spectrum registered in the NIST Mass Spectrometry Database. A compound that is unknown or unregistered can be identified only tentatively, and only by an expert. If a reliable theory can be developed, new compounds will be identified with greater confidence and the analytical power of mass spectrometry will be increased.
Description
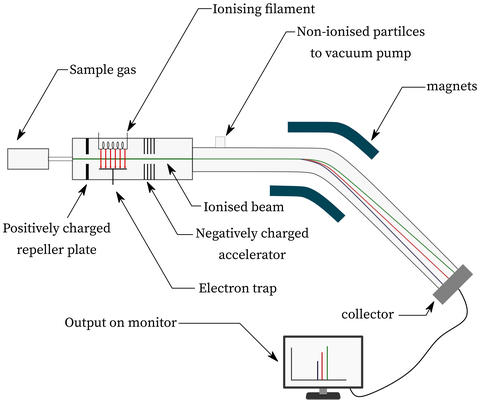
Mass spectrometry separates fragmented molecules on the basis of mass and charge. Research at NIST explores computational methods for modeling fragmentation energetics to enable compound identification.
No Description
Intended Impact
Unknown compounds will be identified more reliably. Quality control for the NIST Mass Spectrometry Database will be enhanced.
Goals
- Develop software tools for predicting microcanonical branching fractions automatically. This is currently too laborious and too technical to be used routinely. (applications: EIMS; MS/MS; CIMS)
- Develop a theoretical approach for predicting the energy deposition function during electron ionization. (application: EIMS)
- Combine the above methods to create software for predicting spectra. (applications: EIMS; MS/MS; CIMS)
RESEARCH ACTIVITIES AND TECHNICAL APPROACH
- Quantum chemistry will be used to determine fragmentation energetics. Reaction pathways will be determined using isopotential searching, heuristics, and other automated methods. Branching fractions will be predicted using RRKM and related theoretical methods.
- New ideas are needed for predicting the energy deposition function; there has been little progress in the past 60 years. One idea is to adapt Kim's BEB theory (developed at NIST) for this purpose.
- Once a successful method is developed, software will be developed and distributed so that the method can be used in mass spectrometry laboratories.
Major Accomplishments
- Thermochemistry may be used as an economical surrogate for kinetics in the formation of sequence ions in peptide MS/MS.
ASSOCIATED PUBLICATIONS
- Irikura, K. K.; Meot-Ner, M; Sieck, L. Wayne; Fant, Andrew D.; Liebman, Joel F., Protonated para-Benzoquinone, J. Org. Chem. 1996, 61, 3167-3171.
- Irikura, K. K.; Johnson, R. D. III, Predicting Unexpected Chemical Reactions by Isopotential Searching, J. Phys. Chem. A 2000, 104, 2191-2194.
- Kim, Y.-K.; Irikura, K. K. Electron-Impact Ionization Cross Sections for Polyatomic Molecules, Radicals, and Ions. In American Institute of Physics Conference Proceedings 543, Atomic and Molecular Data and Their Applications; Berrington, K. A; Bell, K. L., Eds.; American Institute of Physics: College Park, Maryland, 2000; pp. 220-241.
- Haeffner, F.; Merle, J. K.; Irikura, K. K., N-Protonated Isomers as Gateways to Peptide Ion Fragmentation, J. Am. Soc. Mass Spectrom., 2011, 22, 2222-2231.
- Irikura, K. K.; Merle, J. K.; Simón-Manso, Y., Tryptic y++ Fragment Ion Distributions Are Guided by Coulombic Repulsion, J. Am. Soc. Mass Spectrom., 2012, 23, 483-488.
- Irikura, K. K.; Todua, N. G., Facile Smiles-Type Rearrangement in Radical Cations of N-Acyl Arylsulfonamides and Analogs, Rapid Commun. Mass Spectrom., 2014, 28, 829-834.

