Summary
Over the past decade nanoscale science has produced varied and fascinating nanostructures with an extraordinary range of interesting and useful properties. In some instances these nanostructures, from quantum dots to carbon nanotubes, can be used more or less "as is", but in many cases it is necessary to combine disparate species together in precise arrangements to realize the desired functionality. DNA origami has the potential to act as a nanoscale pegboard on which to hang individual nanostructures with molecular precision. The goal of the project is to develop measurement methods that enable the parameters of the assembly process, including speed, yield and precision, as well as their effect on the functionality of the final product, to be understood. Ultimately, this will lead to a set of design rules for processes and structures suitable for reliable nanomanufacturing.
Description
Schematic showing DNA origami in a laser beam
The base pairing of adenine to thymine and guanine to cytosine to form DNA provides a robust molecular recognition scheme that can be used to create a wide variety of well-ordered nanostructures. DNA origami, which uses a long scaffold strand folded together by specific sets of short staple strands, can be used to generate well-controlled DNA shapes, each approximately 100 nm × 100 nm. Selected staple strands can be functionalized with binding moieties, such as complementary DNA for hybridization or biotin, that enable nanostructures such as carbon nanotubes, quantum dots, metal nanoparticles, biomolecules or fluorophores to be attached at predetermined locations.
The flexibility and modularity of this approach make it extremely attractive for making everything from nanoscale optical sensors to targeted drug delivery systems. However, for the potential of this approach to be realized, it is important to understand what the limits are in terms of the speed and yield of this type of process, since these directly affect the ability to scale up production. It is also necessary to understand how different nanostructure attachment strategies and the specifics of the nanostructures being attached can influence the precision that can be achieved in the final product, since this determines how well its functionality can be controlled.
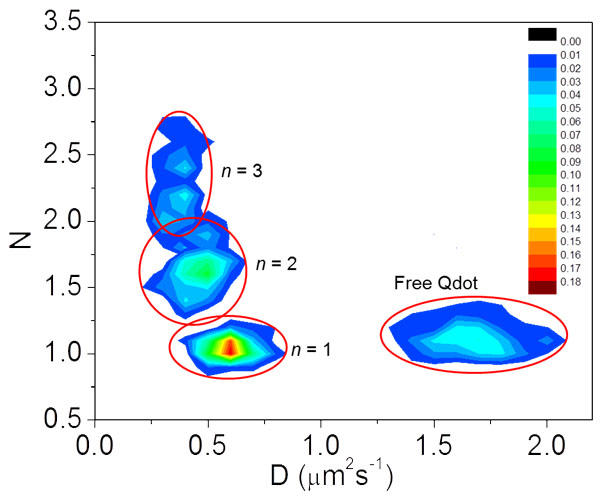
In this project we endeavor to synthesize particular structures in order to understand the issues involved and determine which measurement techniques need to be developed in order to resolve them. We have developed statistical analysis methods to extract forward and backward reaction rate constants from time-course measurements based on atomic force microscope imaging. We have used these measurements to determine how the spacing of binding locations and the number of binding sites available at each location affect the binding of quantum dots to origami. We are using single-particle tracking measurements to follow the progress of the reaction in solution as quantum dots bind to origami with three separate binding sites and are also using of the single-photon counting data acquired in those experiments to determine how the optical response of quantum dot/gold nanoparticle constructs is affected by the number and spacing of the gold particles relative to the quantum dots. Our most recent work involves the development of high-throughput techniques to separate and characterize the products of nanostructure synthesis reactions.
Selected Publications:
Quantum-dot fluorescence lifetime engineering with DNA origami constructs, S. H. Ko, K. Du, and J. A. Liddle, Angewandte Chemie International Edition 52, 1193–1197 (2013).
NIST Publication Database Journal Web Site
Quantum dot-DNA origami binding: a single particle, 3D, real-time tracking study, K. Du, S. H. Ko, G. M. Gallatin, H. P. Yoon, J. A. Liddle, and A. J. Berglund, Chemical Communications 49, 907–909 (2013).
NIST Publication Database Journal Web Site
Nanomanufacturing with DNA origami: factors affecting the kinetics and yield of quantum dot binding, S. H. Ko, G. M. Gallatin, and J. A. Liddle, Advanced Functional Materials 22, 1015-1023 (2012).
NIST Publication Database Journal Web Site

